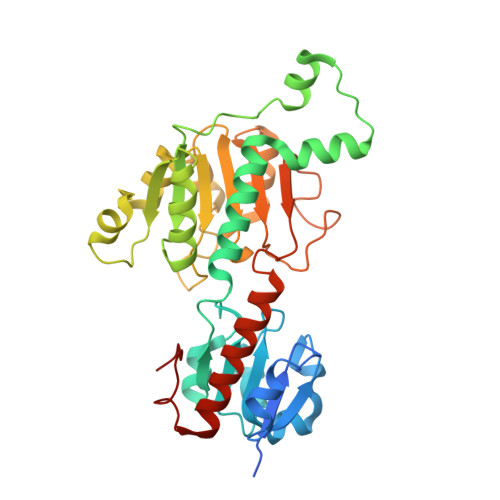
Structure-Guided Design of a High Affinity Inhibitor to Human CtBP.
Hilbert, B.J., Morris, B.L., Ellis, K.C., Paulsen, J.L., Schiffer, C.A., Grossman, S.R., Royer, W.E.(2015) ACS Chem Biol 10: 1118-1127
- PubMed: 25636004 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/cb500820b
- Primary Citation Related Structures:
4U6Q, 4U6S - PubMed Abstract:
Oncogenic transcriptional coregulators C-terminal Binding Protein (CtBP) 1 and 2 possess regulatory d-isomer specific 2-hydroxyacid dehydrogenase (D2-HDH) domains that provide an attractive target for small molecule intervention. Findings that the CtBP substrate 4-methylthio 2-oxobutyric acid (MTOB) can interfere with CtBP oncogenic activity in cell culture and in mice confirm that such inhibitors could have therapeutic benefit. Recent crystal structures of CtBP 1 and 2 revealed that MTOB binds in an active site containing a dominant tryptophan and a hydrophilic cavity, neither of which are present in other D2-HDH family members. Here, we demonstrate the effectiveness of exploiting these active site features for the design of high affinity inhibitors. Crystal structures of two such compounds, phenylpyruvate (PPy) and 2-hydroxyimino-3-phenylpropanoic acid (HIPP), show binding with favorable ring stacking against the CtBP active site tryptophan and alternate modes of stabilizing the carboxylic acid moiety. Moreover, ITC experiments show that HIPP binds to CtBP with an affinity greater than 1000-fold over that of MTOB, and enzymatic assays confirm that HIPP substantially inhibits CtBP catalysis. These results, thus, provide an important step, and additional insights, for the development of highly selective antineoplastic CtBP inhibitors.
- †Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, Massachusetts 01605, United States.
Organizational Affiliation: