Hapten-directed spontaneous disulfide shuffling: a universal technology for site-directed covalent coupling of payloads to antibodies.
Dengl, S., Hoffmann, E., Grote, M., Wagner, C., Mundigl, O., Georges, G., Thorey, I., Stubenrauch, K.G., Bujotzek, A., Josel, H.P., Dziadek, S., Benz, J., Brinkmann, U.(2015) FASEB J 29: 1763-1779
- PubMed: 25670234 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1096/fj.14-263665
- Primary Citation Related Structures:
4S1D - PubMed Abstract:
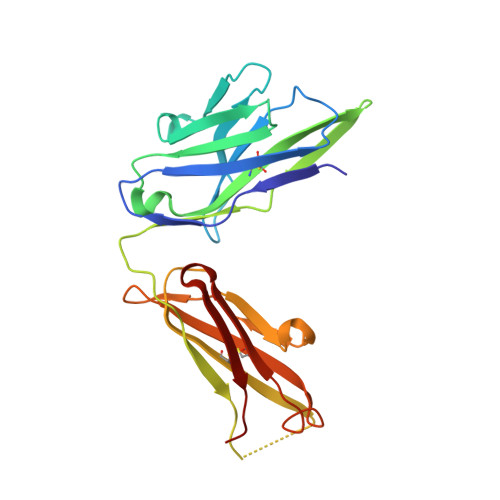
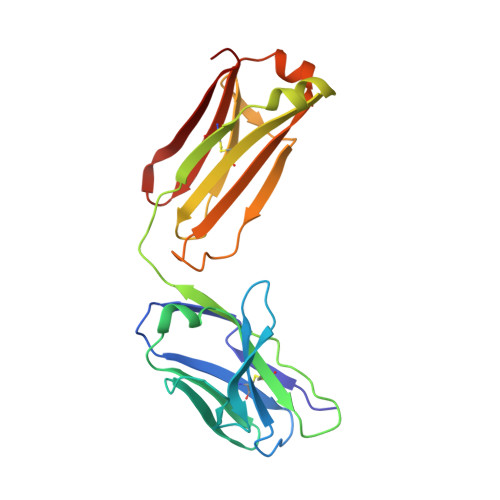
Humanized hapten-binding IgGs were designed with an accessible cysteine close to their binding pockets, for specific covalent payload attachment. Individual analyses of known structures of digoxigenin (Dig)- and fluorescein (Fluo) binding antibodies and a new structure of a biotin (Biot)-binder, revealed a "universal" coupling position (52(+2)) in proximity to binding pockets but without contributing to hapten interactions. Payloads that carry a free thiol are positioned on the antibody and covalently linked to it via disulfides. Covalent coupling is achieved and driven toward complete (95-100%) payload occupancy by spontaneous redox shuffling between antibody and payload. Attachment at the universal position works with different haptens, antibodies, and payloads. Examples are the haptens Fluo, Dig, and Biot combined with various fluorescent or peptidic payloads. Disulfide-bonded covalent antibody-payload complexes do not dissociate in vitro and in vivo. Coupling requires the designed cysteine and matching payload thiol because payload or antibody without the Cys/thiol are not linked (<5% nonspecific coupling). Hapten-mediated positioning is necessary as hapten-thiol-payload is only coupled to antibodies that bind matching haptens. Covalent complexes are more stable in vivo than noncovalent counterparts because digoxigeninylated or biotinylated fluorescent payloads without disulfide-linkage are cleared more rapidly in mice (approximately 50% reduced 48 hour serum levels) compared with their covalently linked counterparts. The coupling technology is applicable to many haptens and hapten binding antibodies (confirmed by automated analyses of the structures of 140 additional hapten binding antibodies) and can be applied to modulate the pharmacokinetics of small compounds or peptides. It is also suitable to link payloads in a reduction-releasable manner to tumor- or tissue-targeting delivery vehicles.
- *Roche Pharma Research & Early Development, Large Molecule Research, Roche Innovation Center Penzberg, Penzberg, Germany; Roche Diagnostics GmbH, Penzberg, Germany; and Roche Discovery Technologies, Roche Innovation Center Basel, Basel, Switzerland ulrich.brinkmann@roche.com stefan.dengl@roche.com eike.hoffmann@roche.com.
Organizational Affiliation: