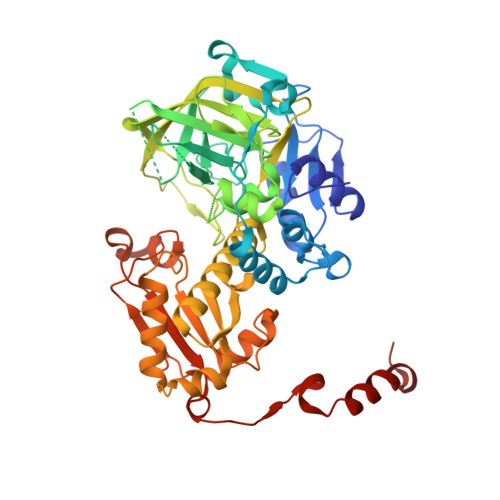
Structure and Mechanism of Ferulic Acid Decarboxylase (FDC1) from Saccharomyces cerevisiae.
Bhuiya, M.W., Lee, S.G., Jez, J.M., Yu, O.(2015) Appl Environ Microbiol 81: 4216-4223
- PubMed: 25862228
- DOI: https://doi.org/10.1128/AEM.00762-15
- Primary Citation of Related Structures:
4S13 - PubMed Abstract:
The nonoxidative decarboxylation of aromatic acids occurs in a range of microbes and is of interest for bioprocessing and metabolic engineering. Although phenolic acid decarboxylases provide useful tools for bioindustrial applications, the molecular bases for how these enzymes function are only beginning to be examined. Here we present the 2.35-Å-resolution X-ray crystal structure of the ferulic acid decarboxylase (FDC1; UbiD) from Saccharomyces cerevisiae. FDC1 shares structural similarity with the UbiD family of enzymes that are involved in ubiquinone biosynthesis. The position of 4-vinylphenol, the product of p-coumaric acid decarboxylation, in the structure identifies a large hydrophobic cavity as the active site. Differences in the β2e-α5 loop of chains in the crystal structure suggest that the conformational flexibility of this loop allows access to the active site. The structure also implicates Glu285 as the general base in the nonoxidative decarboxylation reaction catalyzed by FDC1. Biochemical analysis showed a loss of enzymatic activity in the E285A mutant. Modeling of 3-methoxy-4-hydroxy-5-decaprenylbenzoate, a partial structure of the physiological UbiD substrate, in the binding site suggests that an ∼30-Å-long pocket adjacent to the catalytic site may accommodate the isoprenoid tail of the substrate needed for ubiquinone biosynthesis in yeast. The three-dimensional structure of yeast FDC1 provides a template for guiding protein engineering studies aimed at optimizing the efficiency of aromatic acid decarboxylation reactions in bioindustrial applications.
- Conagen, Inc., Bedford, Massachusetts, USA.
Organizational Affiliation: