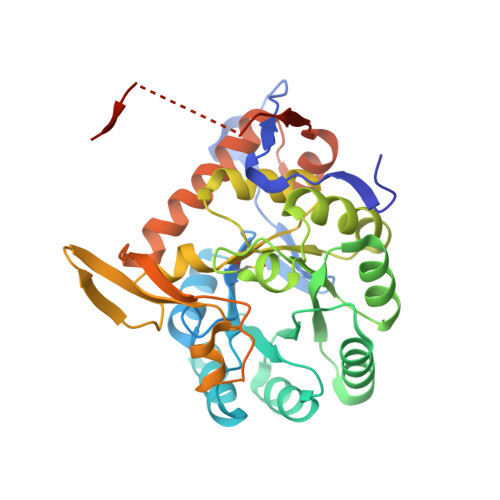
Structure of Cryptosporidium IMP dehydrogenase bound to an inhibitor with in vivo antiparasitic activity.
Kim, Y., Makowska-Grzyska, M., Gorla, S.K., Gollapalli, D.R., Cuny, G.D., Joachimiak, A., Hedstrom, L.(2015) Acta Crystallogr F Struct Biol Commun 71: 531-538
- PubMed: 25945705 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15000187
- Primary Citation Related Structures:
4RV8 - PubMed Abstract:
Inosine 5'-monophosphate dehydrogenase (IMPDH) is a promising target for the treatment of Cryptosporidium infections. Here, the structure of C. parvum IMPDH (CpIMPDH) in complex with inosine 5'-monophosphate (IMP) and P131, an inhibitor with in vivo anticryptosporidial activity, is reported. P131 contains two aromatic groups, one of which interacts with the hypoxanthine ring of IMP, while the second interacts with the aromatic ring of a tyrosine in the adjacent subunit. In addition, the amine and NO2 moieties bind in hydrated cavities, forming water-mediated hydrogen bonds to the protein. The design of compounds to replace these water molecules is a new strategy for the further optimization of C. parvum inhibitors for both antiparasitic and antibacterial applications.
- Center for Structural Genomics of Infectious Diseases, Computational Institute, University of Chicago, 5735 S. Ellis Avenue, Chicago, IL 60637, USA.
Organizational Affiliation: