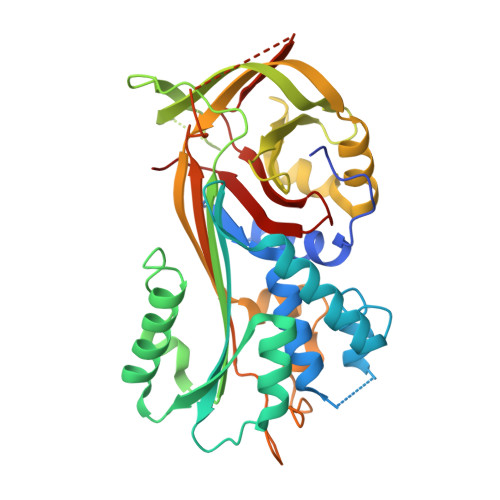
Structural and Inhibitory Effects of Hinge Loop Mutagenesis in Serpin-2 from the Malaria Vector Anopheles gambiae.
Zhang, X., Meekins, D.A., An, C., Zolkiewski, M., Battaile, K.P., Kanost, M.R., Lovell, S., Michel, K.(2015) J Biological Chem 290: 2946-2956
- PubMed: 25525260 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.625665
- Primary Citation Related Structures:
4RO9, 4ROA, 4RSQ - PubMed Abstract:
Serpin-2 (SRPN2) is a key negative regulator of the melanization response in the malaria vector Anopheles gambiae. SRPN2 irreversibly inhibits clip domain serine proteinase 9 (CLIPB9), which functions in a serine proteinase cascade culminating in the activation of prophenoloxidase and melanization. Silencing of SRPN2 in A. gambiae results in spontaneous melanization and decreased life span and is therefore a promising target for vector control. The previously determined structure of SRPN2 revealed a partial insertion of the hinge region of the reactive center loop (RCL) into β sheet A. This partial hinge insertion participates in heparin-linked activation in other serpins, notably antithrombin III. SRPN2 does not contain a heparin binding site, and any possible mechanistic function of the hinge insertion was previously unknown. To investigate the function of the SRPN2 hinge insertion, we developed three SRPN2 variants in which the hinge regions are either constitutively expelled or inserted and analyzed their structure, thermostability, and inhibitory activity. We determined that constitutive hinge expulsion resulted in a 2.7-fold increase in the rate of CLIPB9Xa inhibition, which is significantly lower than previous observations of allosteric serpin activation. Furthermore, we determined that stable insertion of the hinge region did not appreciably decrease the accessibility of the RCL to CLIPB9. Together, these results indicate that the partial hinge insertion in SRPN2 does not participate in the allosteric activation observed in other serpins and instead represents a molecular trade-off between RCL accessibility and efficient formation of an inhibitory complex with the cognate proteinase.
- From the Division of Biology, Kansas State University, Manhattan, Kansas 66506.
Organizational Affiliation: