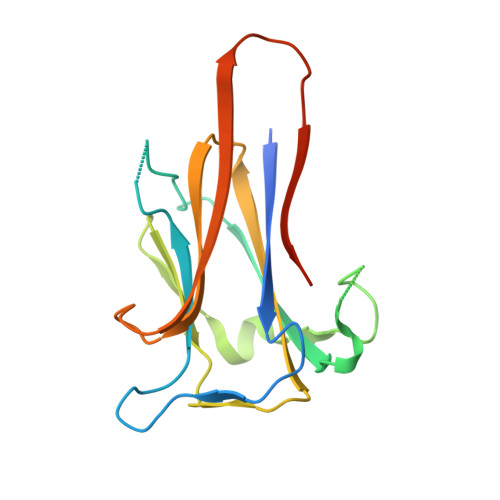
Structure of KRT4 binding domain of Srr-1 from Streptococcus agalactiae reveals a novel beta-sheet complementation.
Sundaresan, R., Samen, U., Ponnuraj, K.(2015) Int J Biol Macromol 75C: 97-105
- PubMed: 25603146
- DOI: https://doi.org/10.1016/j.ijbiomac.2014.12.048
- Primary Citation Related Structures:
4RMB - PubMed Abstract:
The serine rich repeat protein-1 (Srr-1) is an adhesive protein of Streptococcus agalactiae. It is the first bacterial protein identified to interact with human keratin 4 (K4 or KRT4). Within Srr-1, the residues 311-641 constitute the non-repeat ligand binding region (Srr-1-BR(311-641)). The C-terminal part of Srr-1-BR(311-641), comprising of residues 485-642 (termed Srr-1-K4BD), have been identified to bind to K4. Here we report the crystal structure of recombinant Srr-1-K4BD(485-642) and its possible mode of interaction with K4 through docking studies. The dimeric structure of Srr-1-K4BD(485-642) reveals a novel two way "slide lock" parallel β-sheet complementation where the C-terminal strand of one monomer is positioned anti-parallel to the N-terminal strand of the adjacent monomer and this arrangement is not seen so far in any of the homologous structures. The dimerization of Srr-1-K4BD(485-642) observed both in the crystal structure and in solution suggests that similar domain association could also be possible in in vivo and we propose this association would likely generate a new binding site for another host molecule. It is likely that the adhesin can recognize multiple ligands using its ligand binding sub-domains through their intra and inter domain association with one another.
- Centre of Advanced Study in Crystallography and Biophysics, University of Madras, Guindy Campus, Chennai 600 025, India.
Organizational Affiliation: