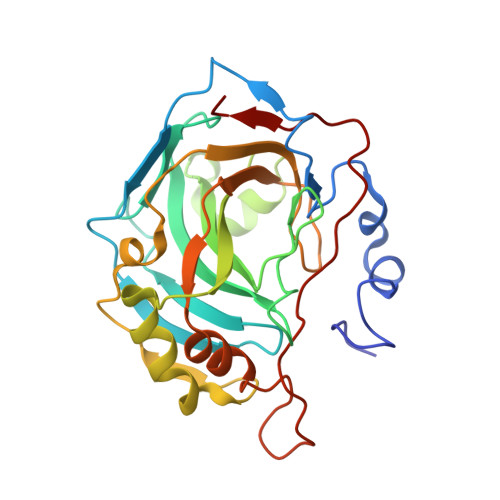
Saccharin: A lead compound for structure-based drug design of carbonic anhydrase IX inhibitors.
Mahon, B.P., Hendon, A.M., Driscoll, J.M., Rankin, G.M., Poulsen, S.A., Supuran, C.T., McKenna, R.(2015) Bioorg Med Chem 23: 849-854
- PubMed: 25614109 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmc.2014.12.030
- Primary Citation Related Structures:
4RIU, 4RIV - PubMed Abstract:
Carbonic anhydrase IX (CA IX) is a key modulator of aggressive tumor behavior and a prognostic marker and target for several cancers. Saccharin (SAC) based compounds may provide an avenue to overcome CA isoform specificity, as they display both nanomolar affinity and preferential binding, for CA IX compared to CA II (>50-fold for SAC and >1000-fold when SAC is conjugated to a carbohydrate moiety). The X-ray crystal structures of SAC and a SAC-carbohydrate conjugate bound to a CA IX-mimic are presented and compared to CA II. The structures provide substantial new insight into the mechanism of SAC selective CA isoform inhibition.
- Department of Biochemistry and Molecular Biology, College of Medicine, University of Florida, Box 100245, Gainesville, FL 32610, USA.
Organizational Affiliation: