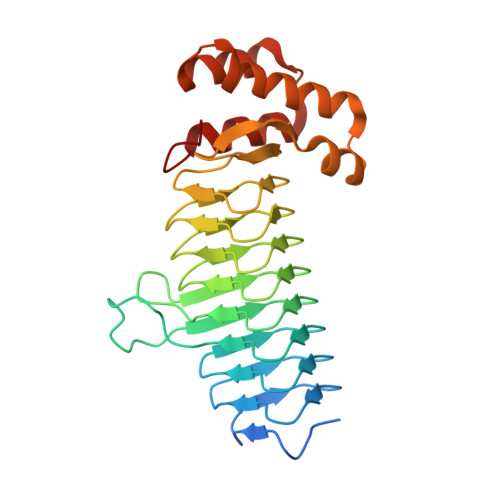
Structures of Bacteroides fragilis uridine 5'-diphosphate-N-acetylglucosamine (UDP-GlcNAc) acyltransferase (BfLpxA).
Ngo, A., Fong, K.T., Cox, D.L., Chen, X., Fisher, A.J.(2015) Acta Crystallogr D Biol Crystallogr 71: 1068-1076
- PubMed: 25945572 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004715003326
- Primary Citation Related Structures:
4R36, 4R37 - PubMed Abstract:
Uridine 5'-diphosphate-N-acetylglucosamine (UDP-GlcNAc) acyltransferase (LpxA) catalyzes a reversible reaction for adding an O-acyl group to the GlcNAc in UDP-GlcNAc in the first step of lipid A biosynthesis. Lipid A constitutes a major component of lipopolysaccharides, also referred to as endotoxins, which form the outer monolayer of the outer membrane of Gram-negative bacteria. Ligand-free and UDP-GlcNAc-bound crystal structures of LpxA from Bacteroides fragilis NCTC 9343, the most common pathogenic bacteria found in abdominal abscesses, have been determined and are presented here. The enzyme crystallizes in a cubic space group, with the crystallographic threefold axis generating the biological functional homotrimer and with each monomer forming a nine-rung left-handed β-helical (LβH) fold in the N-terminus followed by an α-helical motif in the C-terminus. The structure is highly similar to LpxA from other organisms. Yet, despite sharing a similar LβH structure with LpxAs from Escherichia coli and others, previously unseen calcium ions are observed on the threefold axis in B. fragilis LpxA to help stabilize the trimeric assembly.
- Department of Chemistry, University of California, One Shields Avenue, Davis, CA 95616, USA.
Organizational Affiliation: