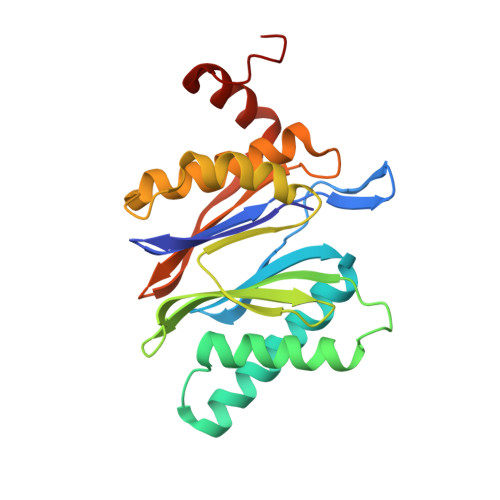
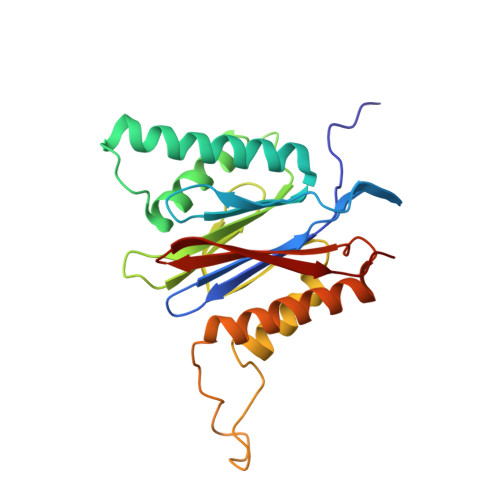
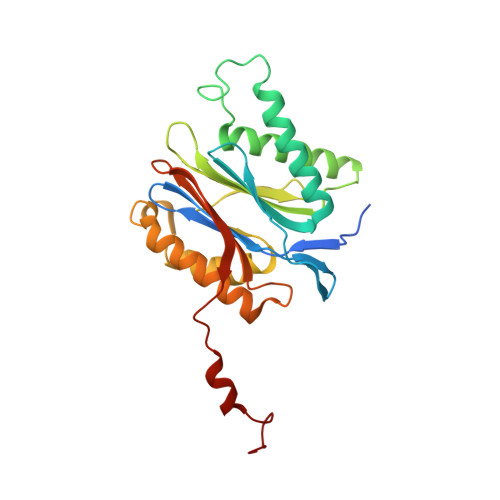
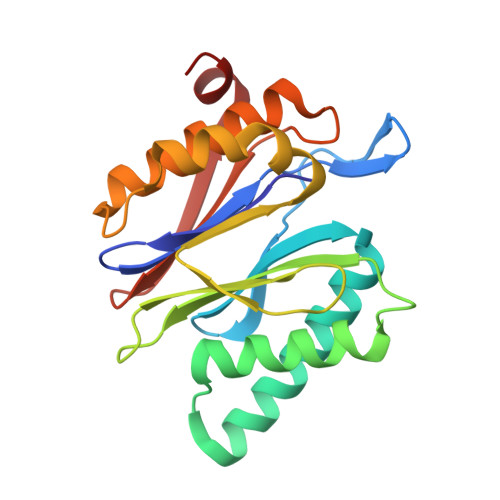
Bortezomib-Resistant Mutant Proteasomes: Structural and Biochemical Evaluation with Carfilzomib and ONX 0914.
Huber, E.M., Heinemeyer, W., Groll, M.(2015) Structure 23: 407-417
- PubMed: 25599643 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2014.11.019
- Primary Citation Related Structures:
4QUX, 4QUY, 4QV0, 4QV1, 4QV3, 4QV4, 4QV5, 4QV6, 4QV7, 4QV8, 4QV9, 4QVL, 4QVM, 4QVN, 4QVP, 4QVQ, 4QVV, 4QVW, 4QVY, 4QW0, 4QW1, 4QW3, 4QW4, 4QW5, 4QW6, 4QW7, 4QWF, 4QWG, 4QWI, 4QWJ, 4QWK, 4QWL, 4QWR, 4QWS, 4QWU, 4QWX, 4QXJ, 4QZ0, 4QZ1, 4QZ2, 4QZ3, 4QZ4, 4QZ5, 4QZ6, 4QZ7, 4QZW, 4QZX, 4QZZ, 4R00 - PubMed Abstract:
Inhibition of the 20S proteasome by bortezomib (Velcade) constitutes a successfully applied therapy for blood cancer. However, emerging resistance restricts its medicinal use. For example, mutations in the proteolytically active β5-subunit of the proteasome, the main target of inhibitors, were reported to impair drug binding and thus to reduce therapeutic efficacy. Using yeast as a model system, we describe here a systematic evaluation of these mutations by cell growth analysis, proteasome inhibition assays, and X-ray crystallography. The 11 mutants examined display decreased proliferation rates, impaired proteolytic activity, and marked resistance to bortezomib as well as the α',β'-epoxyketone inhibitors carfilzomib (Kyprolis) and ONX 0914, while the second-generation compound carfilzomib was the least affected. In total, 49 proteasome X-ray structures, including structural data on proteasome-carfilzomib complexes, reveal three distinct molecular mechanisms that hamper both drug binding and natural substrate turnover to an extent that is still compatible with cell survival.
- Center for Integrated Protein Science at the Department Chemie, Lehrstuhl für Biochemie, Technische Universität München, Lichtenbergstraße 4, 85747 Garching, Germany. Electronic address: eva.huber@mytum.de.
Organizational Affiliation: