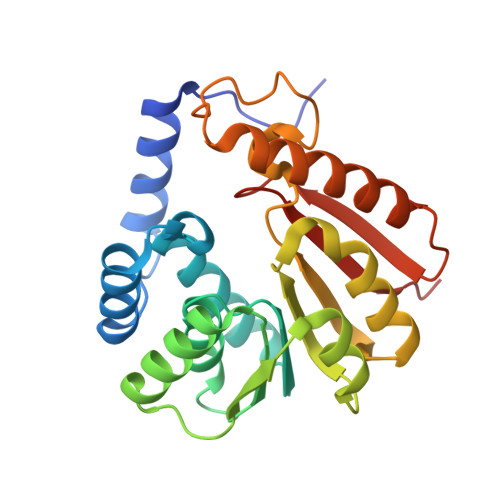
Structure and Biophysical Characterization of the S-Adenosylmethionine-dependent O-Methyltransferase PaMTH1, a Putative Enzyme Accumulating during Senescence of Podospora anserina.
Chatterjee, D., Kudlinzki, D., Linhard, V., Saxena, K., Schieborr, U., Gande, S.L., Wurm, J.P., Wohnert, J., Abele, R., Rogov, V.V., Dotsch, V., Osiewacz, H.D., Sreeramulu, S., Schwalbe, H.(2015) J Biological Chem 290: 16415-16430
- PubMed: 25979334 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.660829
- Primary Citation Related Structures:
4QVK, 4YMG, 4YMH - PubMed Abstract:
Low levels of reactive oxygen species (ROS) act as important signaling molecules, but in excess they can damage biomolecules. ROS regulation is therefore of key importance. Several polyphenols in general and flavonoids in particular have the potential to generate hydroxyl radicals, the most hazardous among all ROS. However, the generation of a hydroxyl radical and subsequent ROS formation can be prevented by methylation of the hydroxyl group of the flavonoids. O-Methylation is performed by O-methyltransferases, members of the S-adenosyl-l-methionine (SAM)-dependent O-methyltransferase superfamily involved in the secondary metabolism of many species across all kingdoms. In the filamentous fungus Podospora anserina, a well established aging model, the O-methyltransferase (PaMTH1) was reported to accumulate in total and mitochondrial protein extracts during aging. In vitro functional studies revealed flavonoids and in particular myricetin as its potential substrate. The molecular architecture of PaMTH1 and the mechanism of the methyl transfer reaction remain unknown. Here, we report the crystal structures of PaMTH1 apoenzyme, PaMTH1-SAM (co-factor), and PaMTH1-S-adenosyl homocysteine (by-product) co-complexes refined to 2.0, 1.9, and 1.9 Å, respectively. PaMTH1 forms a tight dimer through swapping of the N termini. Each monomer adopts the Rossmann fold typical for many SAM-binding methyltransferases. Structural comparisons between different O-methyltransferases reveal a strikingly similar co-factor binding pocket but differences in the substrate binding pocket, indicating specific molecular determinants required for substrate selection. Furthermore, using NMR, mass spectrometry, and site-directed active site mutagenesis, we show that PaMTH1 catalyzes the transfer of the methyl group from SAM to one hydroxyl group of the myricetin in a cation-dependent manner.
- From the Institute for Organic Chemistry and Chemical Biology, Center for Biomolecular Magnetic Resonance (BMRZ), Johann Wolfgang Goethe University, Max-von-Laue-Strasse 7, D-60438 Frankfurt am Main, Germany.
Organizational Affiliation: