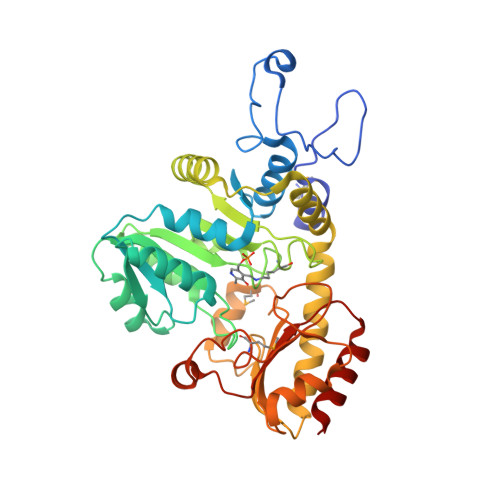
Structural dynamics of a methionine gamma-lyase for calicheamicin biosynthesis: Rotation of the conserved tyrosine stacking with pyridoxal phosphate.
Cao, H., Tan, K., Wang, F., Bigelow, L., Yennamalli, R.M., Jedrzejczak, R., Babnigg, G., Bingman, C.A., Joachimiak, A., Kharel, M.K., Singh, S., Thorson, J.S., Phillips, G.N.(2016) Struct Dyn 3: 034702-034702
- PubMed: 27191010 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1063/1.4948539
- Primary Citation Related Structures:
4Q31 - PubMed Abstract:
CalE6 from Micromonospora echinospora is a (pyridoxal 5' phosphate) PLP-dependent methionine γ-lyase involved in the biosynthesis of calicheamicins. We report the crystal structure of a CalE6 2-(N-morpholino)ethanesulfonic acid complex showing ligand-induced rotation of Tyr100, which stacks with PLP, resembling the corresponding tyrosine rotation of true catalytic intermediates of CalE6 homologs. Elastic network modeling and crystallographic ensemble refinement reveal mobility of the N-terminal loop, which involves both tetrameric assembly and PLP binding. Modeling and comparative structural analysis of PLP-dependent enzymes involved in Cys/Met metabolism shine light on the functional implications of the intrinsic dynamic properties of CalE6 in catalysis and holoenzyme maturation.
- Biosciences at Rice, Rice University , 6100 Main St., Houston, Texas 77005, USA.
Organizational Affiliation: