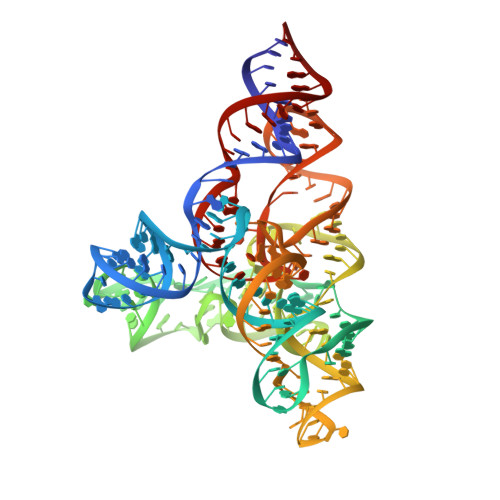
Speciation of a group I intron into a lariat capping ribozyme.
Meyer, M., Nielsen, H., Olieric, V., Roblin, P., Johansen, S.D., Westhof, E., Masquida, B.(2014) Proc Natl Acad Sci U S A 111: 7659-7664
- PubMed: 24821772 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1322248111
- Primary Citation Related Structures:
4P8Z, 4P9R, 6GYV - PubMed Abstract:
The lariat-capping (LC) ribozyme is a natural ribozyme isolated from eukaryotic microorganisms. Despite apparent structural similarity to group I introns, the LC ribozyme catalyzes cleavage by a 2',5' branching reaction, leaving the 3' product with a 3-nt lariat cap that functionally substitutes for a conventional mRNA cap in the downstream pre-mRNA encoding a homing endonuclease. We describe the crystal structures of the precleavage and postcleavage LC ribozymes, which suggest that structural features inherited from group I ribozymes have undergone speciation due to profound changes in molecular selection pressure, ultimately giving rise to an original branching ribozyme family. The structures elucidate the role of key elements that regulate the activity of the LC ribozyme by conformational switching and suggest a mechanism by which the signal for branching is transmitted to the catalytic core. The structures also show how conserved interactions twist residues, forming the lariat to join chemical groups involved in branching.
- Architecture et Réactivité de l'ARN, Unité Propre de Recherche 9002, Institut de Biologie Moléculaire et Cellulaire, Université de Strasbourg, Centre National de la Recherche Scientifique, 67084 Strasbourg, France;
Organizational Affiliation: