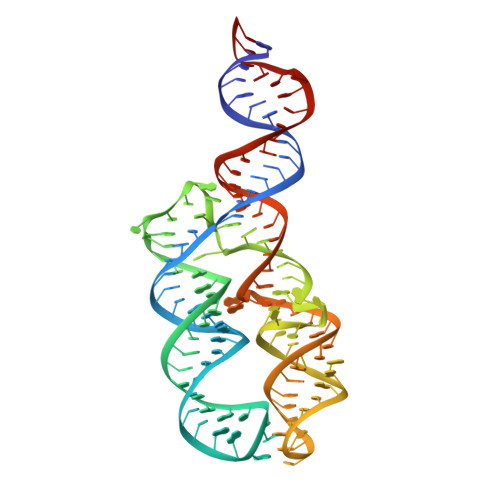
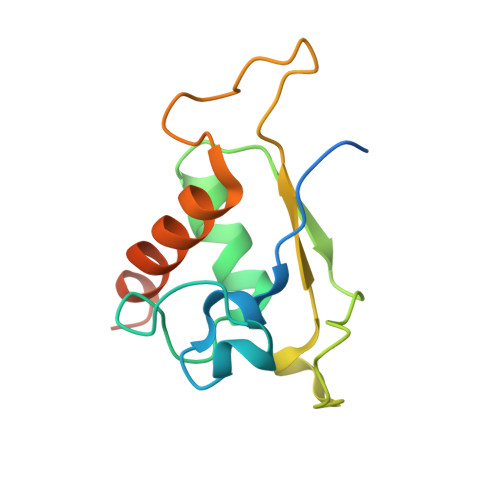
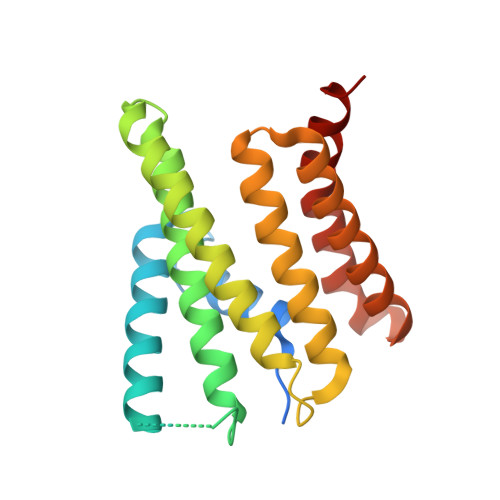
SRP RNA remodeling by SRP68 explains its role in protein translocation.
Grotwinkel, J.T., Wild, K., Segnitz, B., Sinning, I.(2014) Science 344: 101-104
- PubMed: 24700861 Search on PubMed
- DOI: https://doi.org/10.1126/science.1249094
- Primary Citation Related Structures:
4P3E, 4P3F, 4P3G - PubMed Abstract:
The signal recognition particle (SRP) is central to membrane protein targeting; SRP RNA is essential for SRP assembly, elongation arrest, and activation of SRP guanosine triphosphatases. In eukaryotes, SRP function relies on the SRP68-SRP72 heterodimer. We present the crystal structures of the RNA-binding domain of SRP68 (SRP68-RBD) alone and in complex with SRP RNA and SRP19. SRP68-RBD is a tetratricopeptide-like module that binds to a RNA three-way junction, bends the RNA, and inserts an α-helical arginine-rich motif (ARM) into the major groove. The ARM opens the conserved 5f RNA loop, which in ribosome-bound SRP establishes a contact to ribosomal RNA. Our data provide the structural basis for eukaryote-specific, SRP68-driven RNA remodeling required for protein translocation.
- Heidelberg University Biochemistry Center (BZH), INF 328, D-69120 Heidelberg, Germany.
Organizational Affiliation: