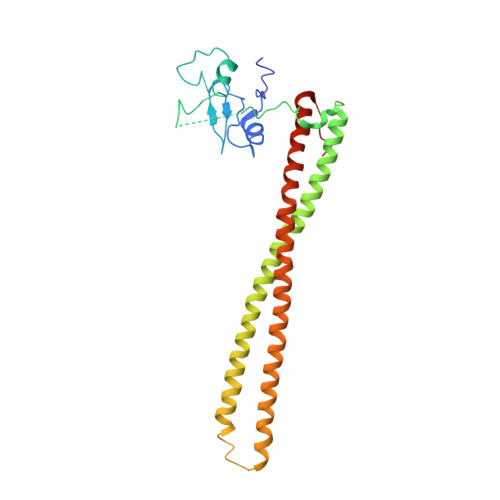
Structural basis of nSH2 regulation and lipid binding in PI3K alpha.
Miller, M.S., Schmidt-Kittler, O., Bolduc, D.M., Brower, E.T., Chaves-Moreira, D., Allaire, M., Kinzler, K.W., Jennings, I.G., Thompson, P.E., Cole, P.A., Amzel, L.M., Vogelstein, B., Gabelli, S.B.(2014) Oncotarget 5: 5198-5208
- PubMed: 25105564 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.18632/oncotarget.2263
- Primary Citation Related Structures:
4OVU, 4OVV - PubMed Abstract:
We report two crystal structures of the wild-type phosphatidylinositol 3-kinase α (PI3Kα) heterodimer refined to 2.9 Å and 3.4 Å resolution: the first as the free enzyme, the second in complex with the lipid substrate, diC4-PIP₂, respectively. The first structure shows key interactions of the N-terminal SH2 domain (nSH2) and iSH2 with the activation loop that suggest a mechanism by which the enzyme is inhibited in its basal state. In the second structure, the lipid substrate binds in a positively charged pocket adjacent to the ATP-binding site, bordered by the P-loop, the activation loop and the iSH2 domain. An additional lipid-binding site was identified at the interface of the ABD, iSH2 and kinase domains. The ability of PI3Kα to bind an additional PIP₂ molecule was confirmed in vitro by fluorescence quenching experiments. The crystal structures reveal key differences in the way the nSH2 domain interacts with wild-type p110α and with the oncogenic mutant p110αH1047R. Increased buried surface area and two unique salt-bridges observed only in the wild-type structure suggest tighter inhibition in the wild-type PI3Kα than in the oncogenic mutant. These differences may be partially responsible for the increased basal lipid kinase activity and increased membrane binding of the oncogenic mutant.
- Medicinal Chemistry, Monash Institute of Pharmaceutical Sciences, Parkville, Victoria, Australia. Present Address: Department of Oncology, Johns Hopkins University School of Medicine, Baltimore Maryland, USA.
Organizational Affiliation: