(Quasi-)Racemic X-ray Structures of Glycosylated and Non-Glycosylated Forms of the Chemokine Ser-CCL1 Prepared by Total Chemical Synthesis.
Okamoto, R., Mandal, K., Sawaya, M.R., Kajihara, Y., Yeates, T.O., Kent, S.B.(2014) Angew Chem Int Ed Engl 53: 5194-5198
- PubMed: 24692304 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201400679
- Primary Citation Related Structures:
4OIJ, 4OIK - PubMed Abstract:
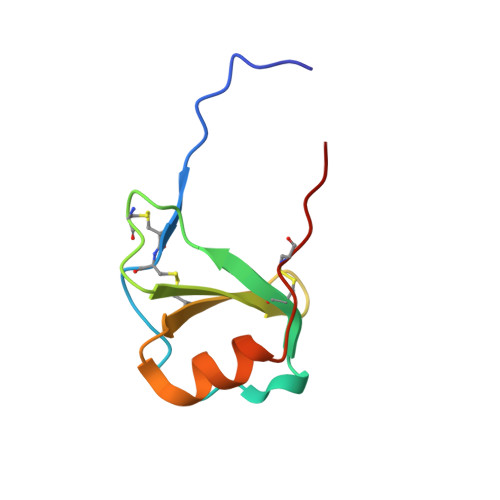
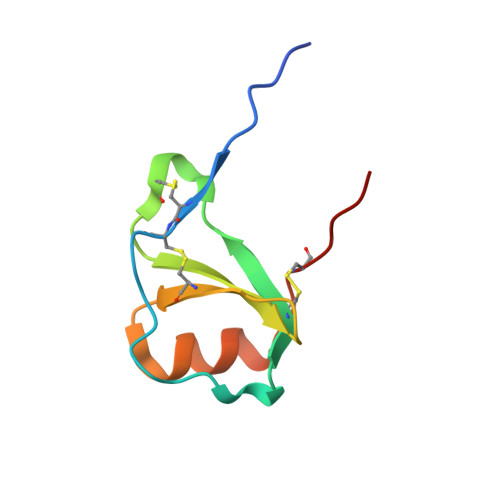
Our goal was to obtain the X-ray crystal structure of the glycosylated chemokine Ser-CCL1. Glycoproteins can be hard to crystallize because of the heterogeneity of the oligosaccharide (glycan) moiety. We used glycosylated Ser-CCL1 that had been prepared by total chemical synthesis as a homogeneous compound containing an N-linked asialo biantennary nonasaccharide glycan moiety of defined covalent structure. Facile crystal formation occurred from a quasi-racemic mixture consisting of glycosylated L-protein and non-glycosylated-D-protein, while no crystals were obtained from the glycosylated L-protein alone. The structure was solved at a resolution of 2.6-2.1 Å. However, the glycan moiety was disordered: only the N-linked GlcNAc sugar was well-defined in the electron density map. A racemic mixture of the protein enantiomers L-Ser-CCL1 and D-Ser-CCL1 was also crystallized, and the structure of the true racemate was solved at a resolution of 2.7-2.15 Å. Superimposition of the structures of the protein moieties of L-Ser-CCL1 and glycosylated-L-Ser-CCL1 revealed there was no significant alteration of the protein structure by N-glycosylation.
- Departments of Chemistry: Biochemistry & Molecular Biology, Institute for Biophysical Dynamics, University of Chicago, Chicago, IL 60637 (USA); Current address: Department of Chemistry, Graduate School of Science, Osaka University, Toyonaka, Osaka, 560-0043, JAPAN.. skent@uchicago.edu, rokamoto@chem.sci.osaka-u.ac.jp, rokamoto@chem.sci.osaka-u.ac.jp.
Organizational Affiliation: