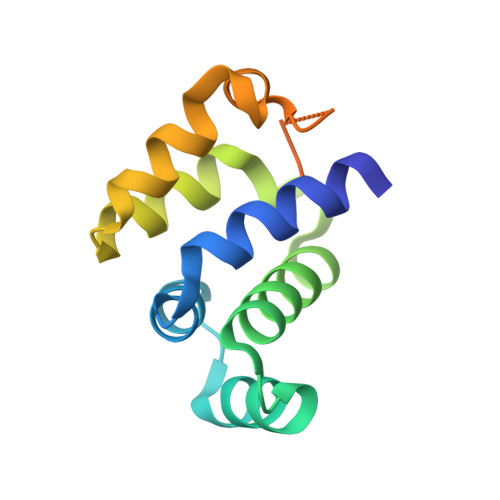
Crystal structure of human Ankyrin G death domain.
Liu, Y., Zhang, Y., Wang, J.H.(2014) Proteins 82: 3476-3482
- PubMed: 25307106 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24702
- Primary Citation Related Structures:
4O6X - PubMed Abstract:
Ankyrins (Ank) are a ubiquitously expressed family of multifunctional membrane adapter proteins. Ankyrin G (AnkG) is critical for assembling and maintenance of the axon initial segment. Here we present the 2.1 Å crystal structure of human AnkG death domain (hAnkG-DD). The core death domain is composed of six α-helices and three 3₁₀-helices. It forms a hydrophobic pocket on the surface of the molecule. The C-terminal tail of the hAnkG-DD curves back to have the aromatic ring of a phenylalanine residue, Phe100 insert into this pocket, which anchors the flexible tail onto the core domain. Related DDs were selected for structure comparison. The major variations are at the C-terminal region, including the α6 and the long C-terminal extension. The results of size exclusion chromatography and analytical ultracentrifugation suggest that hAnkG-DD exists as monomer in solution. Our work should help for the future investigation of the structure-function of AnkG.
- State Key Laboratory of Biomembrane and Membrane Biotechnology, College of Life Sciences, Peking University, Beijing, 100871, China.
Organizational Affiliation: