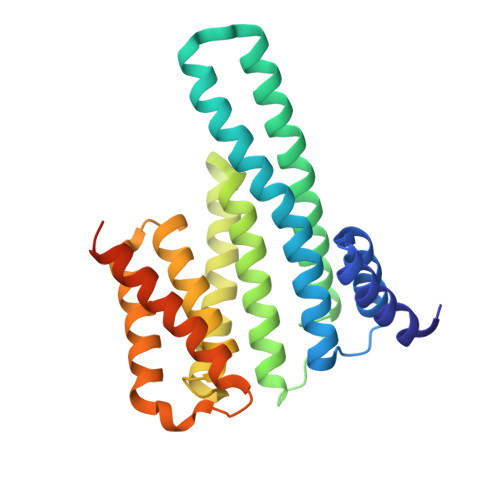
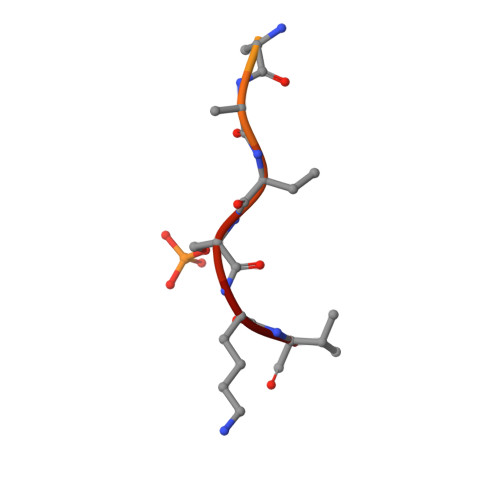
Structural basis for histone mimicry and hijacking of host proteins by influenza virus protein NS1.
Qin, S., Liu, Y., Tempel, W., Eram, M.S., Bian, C., Liu, K., Senisterra, G., Crombet, L., Vedadi, M., Min, J.(2014) Nat Commun 5: 3952-3952
- PubMed: 24853335 Search on PubMed
- DOI: https://doi.org/10.1038/ncomms4952
- Primary Citation Related Structures:
4NW2, 4O42, 4O45, 4O46 - PubMed Abstract:
Pathogens can interfere with vital biological processes of their host by mimicking host proteins. The NS1 protein of the influenza A H3N2 subtype possesses a histone H3K4-like sequence at its carboxyl terminus and has been reported to use this mimic to hijack host proteins. However, this mimic lacks a free N-terminus that is essential for binding to many known H3K4 readers. Here we show that the double chromodomains of CHD1 adopt an 'open pocket' to interact with the free N-terminal amine of H3K4, and the open pocket permits the NS1 mimic to bind in a distinct conformation. We also explored the possibility that NS1 hijacks other cellular proteins and found that the NS1 mimic has access to only a subset of chromatin-associated factors, such as WDR5. Moreover, methylation of the NS1 mimic can not be reversed by the H3K4 demethylase LSD1. Overall, we thus conclude that the NS1 mimic is an imperfect histone mimic.
- 1] Hubei Key Laboratory of Genetic Regulation and Integrative Biology, College of Life Science, Central China Normal University, Wuhan 430079, PR China [2] Structural Genomics Consortium, University of Toronto, 101 College Street, Toronto, Ontario, Canada M5G 1L7 [3].
Organizational Affiliation: