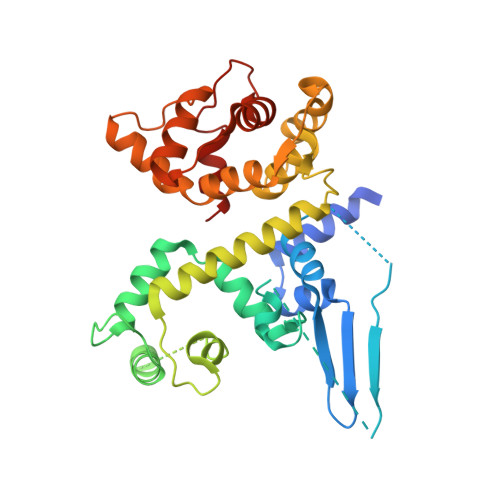
Structural and mechanistic insights into MICU1 regulation of mitochondrial calcium uptake.
Wang, L., Yang, X., Li, S., Wang, Z., Liu, Y., Feng, J., Zhu, Y., Shen, Y.(2014) EMBO J 33: 594-604
- PubMed: 24514027 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/embj.201386523
- Primary Citation Related Structures:
4NSC, 4NSD - PubMed Abstract:
Mitochondrial calcium uptake is a critical event in various cellular activities. Two recently identified proteins, the mitochondrial Ca(2+) uniporter (MCU), which is the pore-forming subunit of a Ca(2+) channel, and mitochondrial calcium uptake 1 (MICU1), which is the regulator of MCU, are essential in this event. However, the molecular mechanism by which MICU1 regulates MCU remains elusive. In this study, we report the crystal structures of Ca(2+)-free and Ca(2+)-bound human MICU1. Our studies reveal that Ca(2+)-free MICU1 forms a hexamer that binds and inhibits MCU. Upon Ca(2+) binding, MICU1 undergoes large conformational changes, resulting in the formation of multiple oligomers to activate MCU. Furthermore, we demonstrate that the affinity of MICU1 for Ca(2+) is approximately 15-20 μM. Collectively, our results provide valuable details to decipher the molecular mechanism of MICU1 regulation of mitochondrial calcium uptake.
- State Key Laboratory of Medicinal Chemical Biology, Nankai University, Tianjin, China.
Organizational Affiliation: