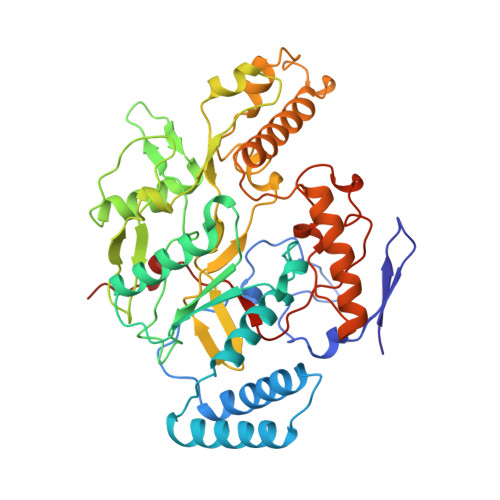
Structural characterization of nitric oxide synthase isoforms reveals striking active-site conservation.
Fischmann, T.O., Hruza, A., Niu, X.D., Fossetta, J.D., Lunn, C.A., Dolphin, E., Prongay, A.J., Reichert, P., Lundell, D.J., Narula, S.K., Weber, P.C.(1999) Nat Struct Biol 6: 233-242
- PubMed: 10074942 Search on PubMed
- DOI: https://doi.org/10.1038/6675
- Primary Citation Related Structures:
3NOS, 4NOS - PubMed Abstract:
Crystal structures of human endothelial nitric oxide synthase (eNOS) and human inducible NOS (iNOS) catalytic domains were solved in complex with the arginine substrate and an inhibitor S-ethylisothiourea (SEITU), respectively. The small molecules bind in a narrow cleft within the larger active-site cavity containing heme and tetrahydrobiopterin. Both are hydrogen-bonded to a conserved glutamate (eNOS E361, iNOS E377). The active-site residues of iNOS and eNOS are nearly identical. Nevertheless, structural comparisons provide a basis for design of isozyme-selective inhibitors. The high-resolution, refined structures of eNOS (2.4 A resolution) and iNOS (2.25 A resolution) reveal an unexpected structural zinc situated at the intermolecular interface and coordinated by four cysteines, two from each monomer.
- Structural Chemistry Department, Schering-Plough Research Institute, Kenilworth, New Jersey 07033, USA. thierry.fischmann@spcorp.com
Organizational Affiliation: