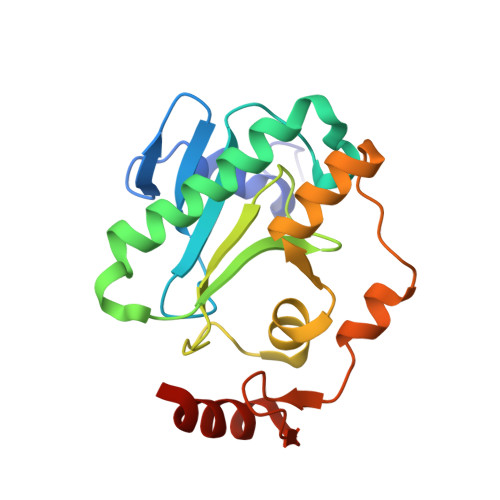
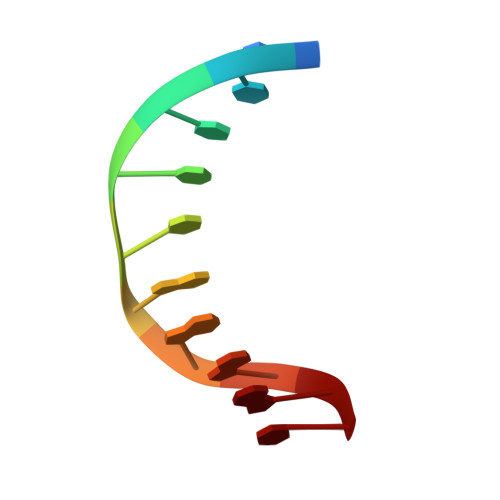
Aprataxin resolves adenylated RNA-DNA junctions to maintain genome integrity.
Tumbale, P., Williams, J.S., Schellenberg, M.J., Kunkel, T.A., Williams, R.S.(2013) Nature 506: 111-115
- PubMed: 24362567 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature12824
- Primary Citation Related Structures:
4NDF, 4NDG, 4NDH, 4NDI - PubMed Abstract:
Faithful maintenance and propagation of eukaryotic genomes is ensured by three-step DNA ligation reactions used by ATP-dependent DNA ligases. Paradoxically, when DNA ligases encounter nicked DNA structures with abnormal DNA termini, DNA ligase catalytic activity can generate and/or exacerbate DNA damage through abortive ligation that produces chemically adducted, toxic 5'-adenylated (5'-AMP) DNA lesions. Aprataxin (APTX) reverses DNA adenylation but the context for deadenylation repair is unclear. Here we examine the importance of APTX to RNase-H2-dependent excision repair (RER) of a lesion that is very frequently introduced into DNA, a ribonucleotide. We show that ligases generate adenylated 5' ends containing a ribose characteristic of RNase H2 incision. APTX efficiently repairs adenylated RNA-DNA, and acting in an RNA-DNA damage response (RDDR), promotes cellular survival and prevents S-phase checkpoint activation in budding yeast undergoing RER. Structure-function studies of human APTX-RNA-DNA-AMP-Zn complexes define a mechanism for detecting and reversing adenylation at RNA-DNA junctions. This involves A-form RNA binding, proper protein folding and conformational changes, all of which are affected by heritable APTX mutations in ataxia with oculomotor apraxia 1. Together, these results indicate that accumulation of adenylated RNA-DNA may contribute to neurological disease.
- 1] Laboratory of Structural Biology, National Institute of Environmental Health Sciences, NIH, DHHS, Research Triangle Park, North Carolina 27709, USA [2].
Organizational Affiliation: