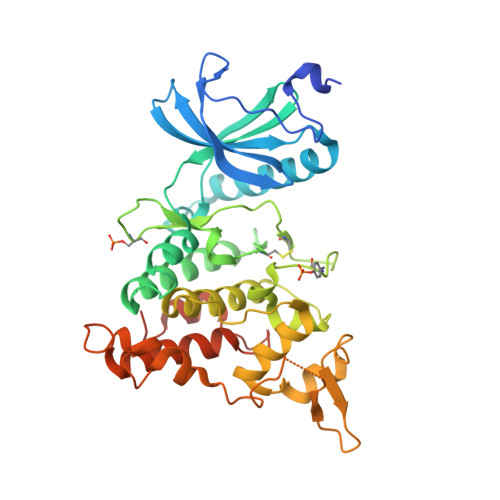
The structure of a dual-specificity tyrosine phosphorylation-regulated kinase 1A-PKC412 complex reveals disulfide-bridge formation with the anomalous catalytic loop HRD(HCD) cysteine.
Alexeeva, M., Aberg, E., Engh, R.A., Rothweiler, U.(2015) Acta Crystallogr D Biol Crystallogr 71: 1207-1215
- PubMed: 25945585 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004715005106
- Primary Citation Related Structures:
4NCT - PubMed Abstract:
Dual-specificity tyrosine phosphorylation-regulated kinase 1A (DYRK1A) is a protein kinase associated with neuronal development and brain physiology. The DYRK kinases are very unusual with respect to the sequence of the catalytic loop, in which the otherwise highly conserved arginine of the HRD motif is replaced by a cysteine. This replacement, along with the proximity of a potential disulfide-bridge partner from the activation segment, implies a potential for redox control of DYRK family activities. Here, the crystal structure of DYRK1A bound to PKC412 is reported, showing the formation of the disulfide bridge and associated conformational changes of the activation loop. The DYRK kinases represent emerging drug targets for several neurological diseases as well as cancer. The observation of distinct activation states may impact strategies for drug targeting. In addition, the characterization of PKC412 binding offers new insights for DYRK inhibitor discovery.
- Department of Chemistry, The Norwegian Structural Biology Centre, The Arctic University of Norway, 9037 Tromsø, Norway.
Organizational Affiliation: