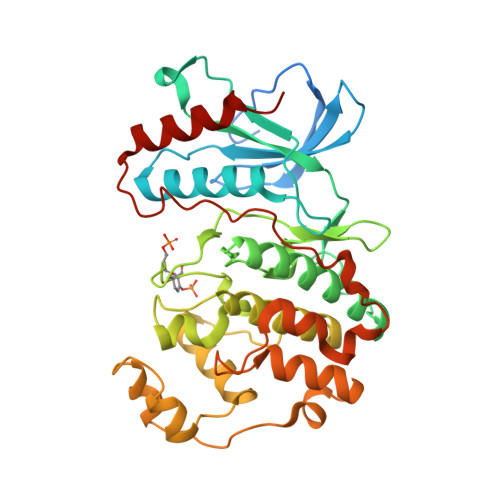
The crystal structure of phosphorylated MAPK13 reveals common structural features and differences in p38 MAPK family activation.
Yurtsever, Z., Scheaffer, S.M., Romero, A.G., Holtzman, M.J., Brett, T.J.(2015) Acta Crystallogr D Biol Crystallogr 71: 790-799
- PubMed: 25849390 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004715001212
- Primary Citation Related Structures:
4MYG - PubMed Abstract:
The p38 MAP kinases (p38 MAPKs) represent an important family centrally involved in mediating extracellular signaling. Recent studies indicate that family members such as MAPK13 (p38δ) display a selective cellular and tissue expression and are therefore involved in specific diseases. Detailed structural studies of all p38 MAPK family members are crucial for the design of specific inhibitors. In order to facilitate such ventures, the structure of MAPK13 was determined in both the inactive (unphosphorylated; MAPK13) and active (dual phosphorylated; MAPK13/pTpY) forms. Here, the first preparation, crystallization and structure determination of MAPK13/pTpY are presented and the structure is compared with the previously reported structure of MAPK13 in order to facilitate studies for structure-based drug design. A comprehensive analysis of inactive versus active structures for the p38 MAPK family is also presented. It is found that MAPK13 undergoes a larger interlobe configurational rearrangement upon activation compared with MAPK14. Surprisingly, the analysis of activated p38 MAPK structures (MAP12/pTpY, MAPK13/pTpY and MAPK14/pTpY) reveals that, despite a high degree of sequence similarity, different side chains are used to coordinate the phosphorylated residues. There are also differences in the rearrangement of the hinge region that occur in MAPK14 compared with MAPK13 which would affect inhibitor binding. A thorough examination of all of the active (phosphorylated) and inactive (unphosphorylated) p38 MAPK family member structures was performed to reveal a common structural basis of activation for the p38 MAP kinase family and to identify structural differences that may be exploited for developing family member-specific inhibitors.
- Biochemistry Program, Washington University School of Medicine, St Louis, MO 63110, USA.
Organizational Affiliation: