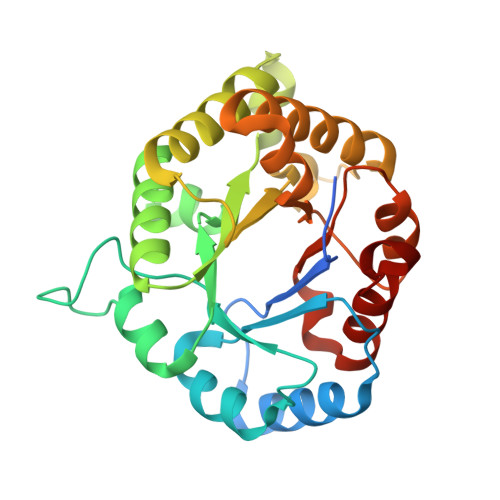
High-Resolution Crystal Structure and Redox Properties of Chloroplastic Triosephosphate Isomerase from Chlamydomonas reinhardtii.
Zaffagnini, M., Michelet, L., Sciabolini, C., Di Giacinto, N., Morisse, S., Marchand, C.H., Trost, P., Fermani, S., Lemaire, S.D.(2014) Mol Plant 7: 101-120
- PubMed: 24157611 Search on PubMed
- DOI: https://doi.org/10.1093/mp/sst139
- Primary Citation Related Structures:
4MKN - PubMed Abstract:
Triosephosphate isomerase (TPI) catalyzes the interconversion of glyceraldehyde-3-phosphate to dihydroxyacetone phosphate. Photosynthetic organisms generally contain two isoforms of TPI located in both cytoplasm and chloroplasts. While the cytoplasmic TPI is involved in the glycolysis, the chloroplastic isoform participates in the Calvin-Benson cycle, a key photosynthetic process responsible for carbon fixation. Compared with its cytoplasmic counterpart, the functional features of chloroplastic TPI have been poorly investigated and its three-dimensional structure has not been solved. Recently, several studies proposed TPI as a potential target of different redox modifications including dithiol/disulfide interchanges, glutathionylation, and nitrosylation. However, neither the effects on protein activity nor the molecular mechanisms underlying these redox modifications have been investigated. Here, we have produced recombinantly and purified TPI from the unicellular green alga Chlamydomonas reinhardtii (Cr). The biochemical properties of the enzyme were delineated and its crystallographic structure was determined at a resolution of 1.1 Å. CrTPI is a homodimer with subunits containing the typical (β/α)8-barrel fold. Although no evidence for TRX regulation was obtained, CrTPI was found to undergo glutathionylation by oxidized glutathione and trans-nitrosylation by nitrosoglutathione, confirming its sensitivity to multiple redox modifications.
- Laboratoire de Biologie Moléculaire et Cellulaire des Eucaryotes, FRE3354 Centre National de la Recherche Scientifique, Université Pierre et Marie Curie, Institut de Biologie Physico-Chimique, 13 rue Pierre et Marie Curie, 75005 Paris, France.
Organizational Affiliation: