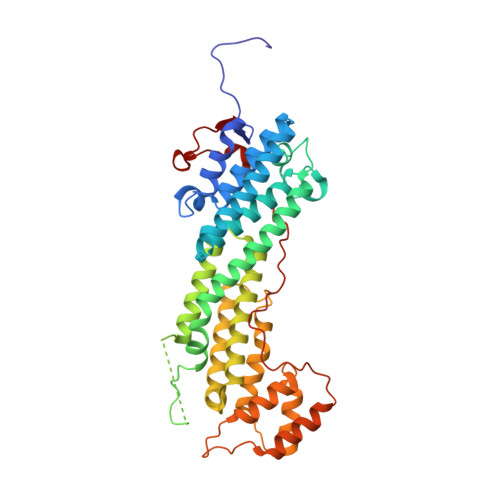
Structural insights into the globular tails of the human type v myosins myo5a, myo5b, and myo5c.
Velvarska, H., Niessing, D.(2013) PLoS One 8: e82065-e82065
- PubMed: 24339992 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0082065
- Primary Citation Related Structures:
4LLI, 4LNZ - PubMed Abstract:
Vertebrate type V myosins (MyoV) Myo5a, Myo5b, and Myo5c mediate transport of several different cargoes. All MyoV paralogs bind to cargo complexes mainly by their C-terminal globular domains. In absence of cargo, the globular domain of Myo5a inhibits its motor domain. Here, we report low-resolution SAXS models for the globular domains from human Myo5a, Myo5b, and Myo5c, which suggest very similar overall shapes of all three paralogs. We determined the crystal structures of globular domains from Myo5a and Myo5b, and provide a homology model for human Myo5c. When we docked the Myo5a crystal structure into a previously published electron microscopy density of the autoinhibited full-length Myo5a, only one domain orientation resulted in a good fit. This structural arrangement suggests the participation of additional region of the globular domain in autoinhibition. Quantification of the interaction of the Myo5a globular domain with its motor complex revealed a tight binding with dissociation half-life in the order of minutes, suggesting a rather slow transition between the active and inactive states.
- Institute of Structural Biology; Helmholtz Zentrum München - German Research Center for Environmental Health, Neuherberg, Germany ; Gene Center and Department of Biochemistry, Ludwig-Maximilians-University, München, Germany.
Organizational Affiliation: