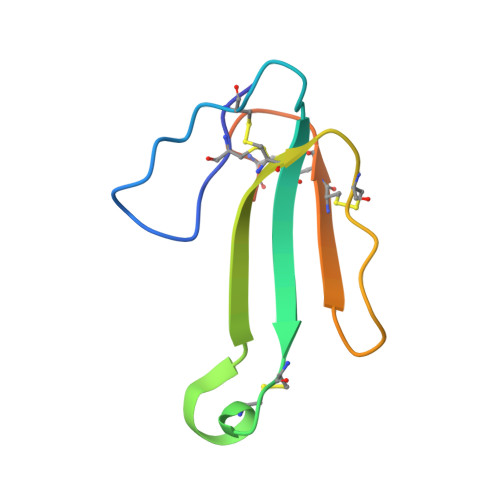
Isolation and Structural and Pharmacological Characterization of alpha-Elapitoxin-Dpp2d, an Amidated Three Finger Toxin from Black Mamba Venom.
Wang, C.I., Reeks, T., Vetter, I., Vergara, I., Kovtun, O., Lewis, R.J., Alewood, P.F., Durek, T.(2014) Biochemistry 53: 3758-3766
- PubMed: 24867092
- DOI: https://doi.org/10.1021/bi5004475
- Primary Citation Related Structures:
4LFT - PubMed Abstract:
We isolated a novel, atypical long-chain three-finger toxin (TFT), α-elapitoxin-Dpp2d (α-EPTX-Dpp2d), from black mamba (Dendroaspis polylepis polylepis) venom. Proteolytic digestion with trypsin and V8 protease, together with MS/MS de novo sequencing, indicated that the mature toxin has an amidated C-terminal arginine, a posttranslational modification rarely observed for snake TFTs. α-EPTX-Dpp2d was found to potently inhibit α7 neuronal nicotinic acetylcholine receptors (nAChR; IC₅₀, 58 ± 24 nM) and muscle-type nAChR (IC₅₀, 114 ± 37 nM) but did not affect α3β2 and α3β4 nAChR isoforms at 1 μM concentrations. Competitive radioligand binding assays demonstrated that α-EPTX-Dpp2d competes with epibatidine binding to the Lymnea stagnalis acetylcholine-binding protein (Ls-AChBP; IC₅₀, 4.9 ± 2.3 nM). The activity profile and binding data are reminiscent of classical long-chain TFTs with a free carboxyl termini, suggesting that amidation does not significantly affect toxin selectivity. The crystal structure of α-EPTX-Dpp2d was determined at 1.7 Å resolution and displayed a dimeric toxin assembly with each monomer positioned in an antiparallel orientation. The dimeric structure is stabilized by extensive intermolecular hydrogen bonds and electrostatic interactions, which raised the possibility that the toxin may exist as a noncovalent homodimer in solution. However, chemical cross-linking and size-exclusion chromatography coupled with multiangle laser light scattering (MALLS) data indicated that the toxin is predominantly monomeric under physiological conditions. Because of its high potency and selectivity, we expect this toxin to be a valuable pharmacological tool for studying the structure and function of nAChRs.
- Division of Chemistry and Structural Biology, Institute for Molecular Bioscience, The University of Queensland, St Lucia , Brisbane, Queensland 4072, Australia.
Organizational Affiliation: