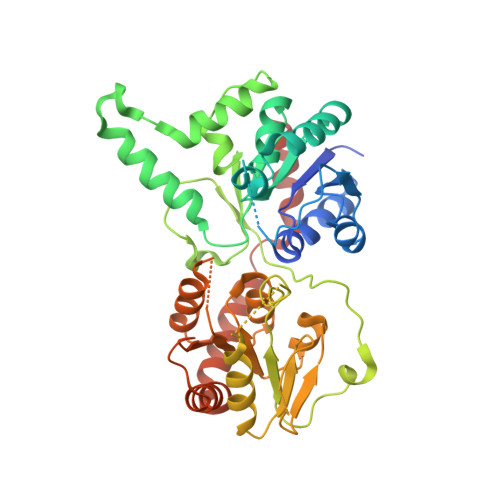
Structural studies of the spinosyn forosaminyltransferase, SpnP.
Isiorho, E.A., Jeon, B.S., Kim, N.H., Liu, H.W., Keatinge-Clay, A.T.(2014) Biochemistry 53: 4292-4301
- PubMed: 24945604 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi5003629
- Primary Citation Related Structures:
4LEI - PubMed Abstract:
Spinosyns A and D (spinosad) are complex polyketide natural products biosynthesized through the cooperation of a modular polyketide synthase and several tailoring enzymes. SpnP catalyzes the final tailoring step, transferring forosamine from a TDP-D-forosamine donor substrate to a spinosyn pseudoaglycone acceptor substrate. Sequence analysis indicated that SpnP belongs to a small group of glycosyltransferases (GTs) that require an auxiliary protein for activation. However, unlike other GTs in this subgroup, no putative auxiliary protein gene could be located in the biosynthetic gene cluster. To learn more about SpnP, the structures of SpnP and its complex with TDP were determined to 2.50 and 3.15 Å resolution, respectively. Binding of TDP causes the reordering of several residues in the donor substrate pocket. SpnP possesses a structural feature that has only been previously observed in the related glycosyltransferase EryCIII, in which it mediates association with the auxiliary protein EryCII. This motif, H-X-R-X5-D-X5-R-X12-20-D-P-X3-W-L-X12-18-E-X4-G, may be predictive of glycosyltransferases that interact with an auxiliary protein. A reverse glycosyl transfer assay demonstrated that SpnP possesses measurable activity in the absence of an auxiliary protein. Our data suggest that SpnP can bind its donor substrate by itself but that the glycosyl transfer reaction is facilitated by an auxiliary protein that aids in the correct folding of a flexible loop surrounding the pseudoaglycone acceptor substrate-binding pocket.
- Division of Medicinal Chemistry, College of Pharmacy, and ‡Department of Chemistry and Biochemistry, The University of Texas at Austin , Austin, Texas 78712, United States.
Organizational Affiliation: