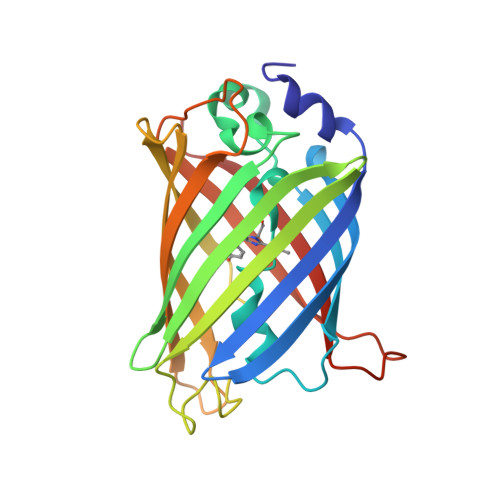
Structural basis for a hand-like site in the calcium sensor CatchER with fast kinetics.
Zhang, Y., Reddish, F., Tang, S., Zhuo, Y., Wang, Y.F., Yang, J.J., Weber, I.T.(2013) Acta Crystallogr D Biol Crystallogr 69: 2309-2319
- PubMed: 24311573 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444913021306
- Primary Citation Related Structures:
4L12, 4L13, 4L1I - PubMed Abstract:
Calcium ions, which are important signaling molecules, can be detected in the endoplasmic reticulum by an engineered mutant of green fluorescent protein (GFP) designated CatchER with a fast off-rate. High resolution (1.78-1.20 Å) crystal structures were analyzed for CatchER in the apo form and in complexes with calcium or gadolinium to probe the binding site for metal ions. While CatchER exhibits a 1:1 binding stoichiometry in solution, two positions were observed for each of the metal ions bound within the hand-like site formed by the carboxylate side chains of the mutated residues S147E, S202D, Q204E, F223E and T225E that may be responsible for its fast kinetic properties. Comparison of the structures of CatchER, wild-type GFP and enhanced GFP confirmed that different conformations of Thr203 and Glu222 are associated with the two forms of Tyr66 of the chromophore which are responsible for the absorbance wavelengths of the different proteins. Calcium binding to CatchER may shift the equilibrium for conformational population of the Glu222 side chain and lead to further changes in its optical properties.
- Department of Chemistry, Georgia State University, Atlanta, GA 30303, USA.
Organizational Affiliation: