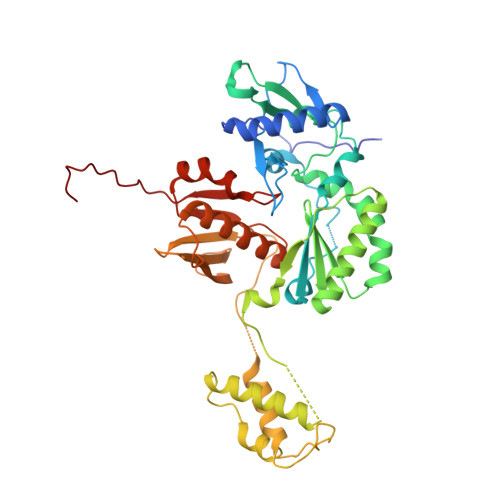
Selective unfolding of one Ribonuclease H domain of HIV reverse transcriptase is linked to homodimer formation.
Zheng, X., Pedersen, L.C., Gabel, S.A., Mueller, G.A., Cuneo, M.J., DeRose, E.F., Krahn, J.M., London, R.E.(2014) Nucleic Acids Res 42: 5361-5377
- PubMed: 24574528 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku143
- Primary Citation Related Structures:
4KSE - PubMed Abstract:
HIV-1 reverse transcriptase (RT), a critical enzyme of the HIV life cycle and an important drug target, undergoes complex and largely uncharacterized conformational rearrangements that underlie its asymmetric folding, dimerization and subunit-selective ribonuclease H domain (RH) proteolysis. In the present article we have used a combination of NMR spectroscopy, small angle X-ray scattering and X-ray crystallography to characterize the p51 and p66 monomers and the conformational maturation of the p66/p66' homodimer. The p66 monomer exists as a loosely structured molecule in which the fingers/palm/connection, thumb and RH substructures are connected by flexible (disordered) linking segments. The initially observed homodimer is asymmetric and includes two fully folded RH domains, while exhibiting other conformational features similar to that of the RT heterodimer. The RH' domain of the p66' subunit undergoes selective unfolding with time constant ∼6.5 h, consistent with destabilization due to residue transfer to the polymerase' domain on the p66' subunit. A simultaneous increase in the intensity of resonances near the random coil positions is characterized by a similar time constant. Consistent with the residue transfer hypothesis, a construct of the isolated RH domain lacking the two N-terminal residues is shown to exhibit reduced stability. These results demonstrate that RH' unfolding is coupled to homodimer formation.
- Laboratory of Structural Biology, NIEHS, National Institutes of Health, Research Triangle Park, NC 27709, USA.
Organizational Affiliation: