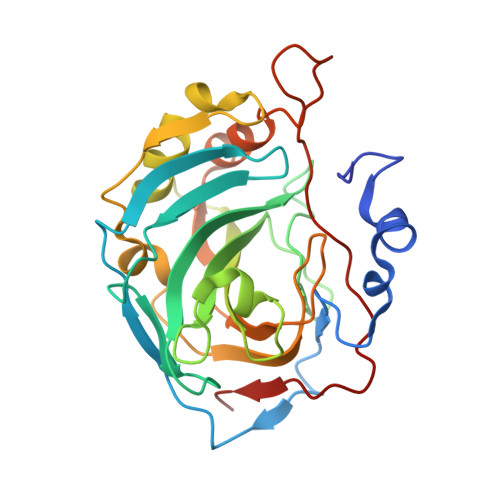
Benzenesulfonamides with pyrimidine moiety as inhibitors of human carbonic anhydrases I, II, VI, VII, XII, and XIII
Capkauskaite, E., Zubriene, A., Smirnov, A., Torresan, J., Kisonaite, M., Kazokaite, J., Gylyte, J., Michailoviene, V., Jogaite, V., Manakova, E., Grazulis, S., Tumkevicius, S., Matulis, D.(2013) Bioorg Med Chem 21: 6937-6947
- PubMed: 24103428 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2013.09.029
- Primary Citation Related Structures:
4KNI, 4KNJ, 4KNM, 4KNN, 4KP5, 4KP8 - PubMed Abstract:
Two groups of benzenesulfonamide derivatives, bearing pyrimidine moieties, were designed and synthesized as inhibitors of carbonic anhydrases (CA). Their binding affinities to six recombinant human CA isoforms I, II, VI, VII, XII, and XIII were determined by the thermal shift assay (TSA). The binding of several inhibitors was measured by isothermal titration calorimetry (ITC). Direct demonstration of compound inhibition was achieved by determining the inhibition constant by stopped-flow CO2 hydration assay. The most potent compounds demonstrated selectivity towards isoform I and affinities of 0.5 nM. The crystal structures of selected compounds in complex with CA II, XII, and XIII were determined to atomic resolution. Compounds described here were compared with previously published pyrimidinebenzenesulfonamides.(1) Systematic structure-activity analysis of 40 compound interactions with six isoforms yields clues for the design of compounds with greater affinities and selectivities towards target CA isoforms.
- Department of Biothermodynamics and Drug Design, Institute of Biotechnology, Vilnius University, Graičiūno 8, Vilnius LT-02241, Lithuania.
Organizational Affiliation: