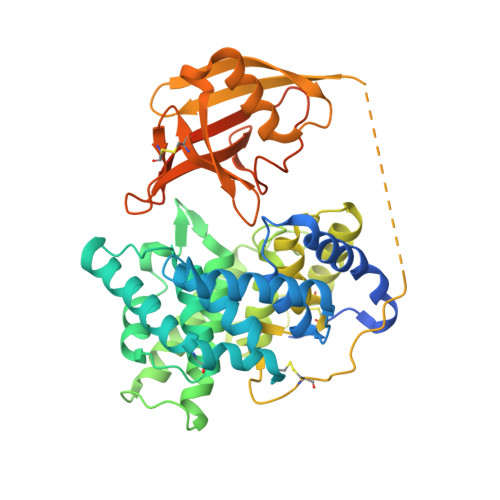
Structural basis for universal corrinoid recognition by the cobalamin transport protein haptocorrin.
Furger, E., Frei, D.C., Schibli, R., Fischer, E., Prota, A.E.(2013) J Biological Chem 288: 25466-25476
- PubMed: 23846701 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.483271
- Primary Citation Related Structures:
4KKI, 4KKJ - PubMed Abstract:
Cobalamin (Cbl; vitamin B12) is an essential micronutrient synthesized only by bacteria. Mammals have developed a sophisticated uptake system to capture the vitamin from the diet. Cbl transport is mediated by three transport proteins: transcobalamin, intrinsic factor, and haptocorrin (HC). All three proteins have a similar overall structure but a different selectivity for corrinoids. Here, we present the crystal structures of human HC in complex with cyanocobalamin and cobinamide at 2.35 and 3.0 Å resolution, respectively. The structures reveal that many of the interactions with the corrin ring are conserved among the human Cbl transporters. However, the non-conserved residues Asn-120, Arg-357, and Asn-373 form distinct interactions allowing for stabilization of corrinoids other than Cbl. A central binding motif forms interactions with the e- and f-side chains of the corrin ring and is conserved in corrinoid-binding proteins of other species. In addition, the α- and β-domains of HC form several unique interdomain contacts and have a higher shape complementarity than those of intrinsic factor and transcobalamin. The stabilization of ligands by all of these interactions is reflected in higher melting temperatures of the protein-ligand complexes. Our structural analysis offers fundamental insights into the unique binding behavior of HC and completes the picture of Cbl interaction with its three transport proteins.
- From the Center for Radiopharmaceutical Sciences and.
Organizational Affiliation: