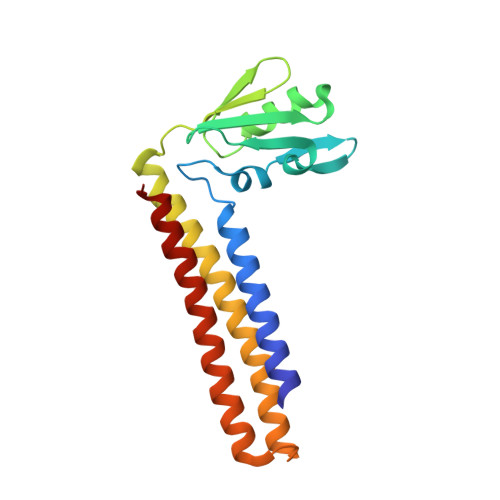
Molecular flexibility of Mycobacterium tuberculosis ribosome recycling factor and its functional consequences: an exploration involving mutants.
Selvaraj, M., Govindan, A., Seshadri, A., Dubey, B., Varshney, U., Vijayan, M.(2013) J Biosci 38: 845-855
- PubMed: 24296887
- DOI: https://doi.org/10.1007/s12038-013-9381-0
- Primary Citation Related Structures:
4KAW, 4KB2, 4KB4, 4KC6, 4KDD - PubMed Abstract:
Internal mobility of the two domain molecule of ribosome recycling factor (RRF) is known to be important for its action. Mycobacterium tuberculosis RRF does not complement E. coli for its deficiency of RRF (in the presence of E. coli EF-G alone). Crystal structure had revealed higher rigidity of the M. tuberculosis RRF due to the presence of additional salt bridges between domains. Two inter-domain salt bridges and one between the linker region and the domain containing C-terminal residues were disrupted by appropriate mutations. Except for a C-terminal deletion mutant, all mutants showed RRF activity in E. coli when M. tuberculosis EF-G was also co-expressed. The crystal structures of the point mutants, that of the C-terminal deletion mutant and that of the protein grown in the presence of a detergent, were determined. The increased mobility resulting from the disruption of the salt bridge involving the hinge region allows the appropriate mutant to weakly complement E. coli for its deficiency of RRF even in the absence of simultaneous expression of the mycobacterial EF-G. The loss of activity of the C-terminal deletion mutant appears to be partly due to the rigidification of the molecule consequent to changes in the hinge region.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India.
Organizational Affiliation: