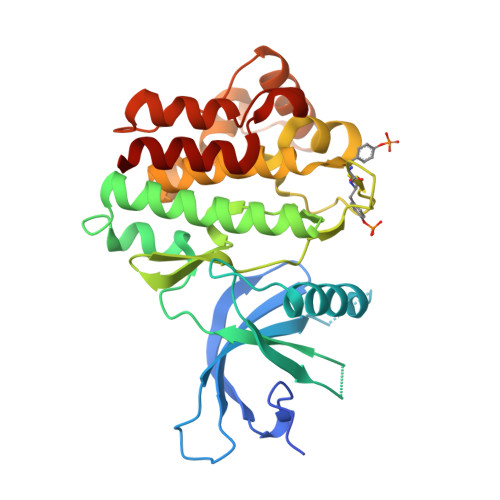
Design and evaluation of novel 8-oxo-pyridopyrimidine Jak1/2 inhibitors.
Labadie, S., Barrett, K., Blair, W.S., Chang, C., Deshmukh, G., Eigenbrot, C., Gibbons, P., Johnson, A., Kenny, J.R., Kohli, P.B., Liimatta, M., Lupardus, P.J., Shia, S., Steffek, M., Ubhayakar, S., Abbema, A.V., Zak, M.(2013) Bioorg Med Chem Lett 23: 5923-5930
- PubMed: 24042009 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2013.08.082
- Primary Citation Related Structures:
4K6Z, 4K77 - PubMed Abstract:
A highly ligand efficient, novel 8-oxo-pyridopyrimidine containing inhibitor of Jak1 and Jak2 isoforms with a pyridone moiety as the hinge-binding motif was discovered. Structure-based design strategies were applied to significantly improve enzyme potency and the polarity of the molecule was adjusted to gain cellular activity. The crystal structures of two representative inhibitors bound to Jak1 were obtained to enable SAR exploration.
- Genentech Inc., 1 DNA Way, South San Francisco, CA 94080, USA. Electronic address: sharadal@gene.com.
Organizational Affiliation: