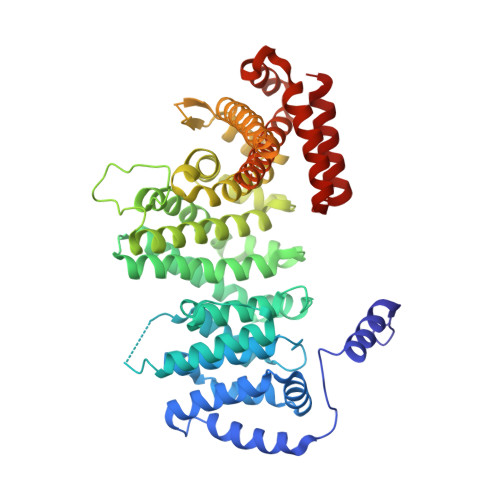
Structure of the human cohesin inhibitor Wapl.
Ouyang, Z., Zheng, G., Song, J., Borek, D.M., Otwinowski, Z., Brautigam, C.A., Tomchick, D.R., Rankin, S., Yu, H.(2013) Proc Natl Acad Sci U S A 110: 11355-11360
- PubMed: 23776203 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1304594110
- Primary Citation Related Structures:
4K6J - PubMed Abstract:
Cohesin, along with positive regulators, establishes sister-chromatid cohesion by forming a ring to circle chromatin. The wings apart-like protein (Wapl) is a key negative regulator of cohesin and forms a complex with precocious dissociation of sisters protein 5 (Pds5) to promote cohesin release from chromatin. Here we report the crystal structure and functional characterization of human Wapl. Wapl contains a flexible, variable N-terminal region (Wapl-N) and a conserved C-terminal domain (Wapl-C) consisting of eight HEAT (Huntingtin, Elongation factor 3, A subunit, and target of rapamycin) repeats. Wapl-C folds into an elongated structure with two lobes. Structure-based mutagenesis maps the functional surface of Wapl-C to two distinct patches (I and II) on the N lobe and a localized patch (III) on the C lobe. Mutating critical patch I residues weaken Wapl binding to cohesin and diminish sister-chromatid resolution and cohesin release from mitotic chromosomes in human cells and Xenopus egg extracts. Surprisingly, patch III on the C lobe does not contribute to Wapl binding to cohesin or its known regulators. Although patch I mutations reduce Wapl binding to intact cohesin, they do not affect Wapl-Pds5 binding to the cohesin subcomplex of sister chromatid cohesion protein 1 (Scc1) and stromal antigen 2 (SA2) in vitro, which is instead mediated by Wapl-N. Thus, Wapl-N forms extensive interactions with Pds5 and Scc1-SA2. Wapl-C interacts with other cohesin subunits and possibly unknown effectors to trigger cohesin release from chromatin.
- Howard Hughes Medical Institute, Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: