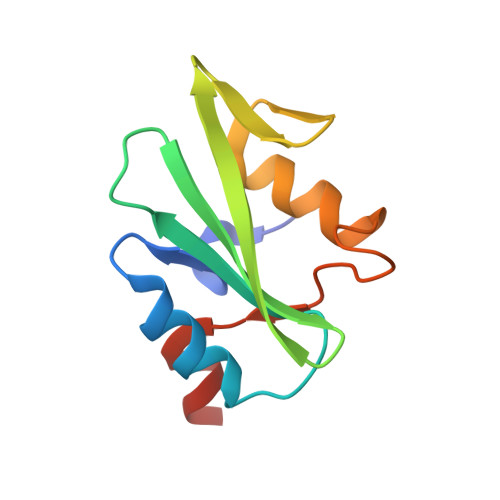
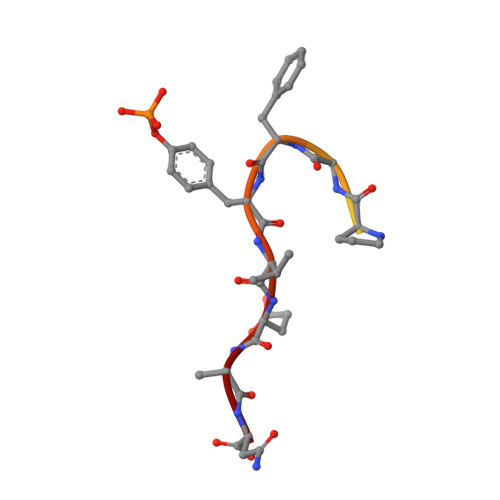
Autoinhibition and Phosphorylation-Induced Activation of Phospholipase C-gamma Isozymes.
Hajicek, N., Charpentier, T.H., Rush, J.R., Harden, T.K., Sondek, J.(2013) Biochemistry 52: 4810-4819
- PubMed: 23777354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi400433b
- Primary Citation Related Structures:
4K44, 4K45 - PubMed Abstract:
Multiple extracellular stimuli, such as growth factors and antigens, initiate signaling cascades through tyrosine phosphorylation and activation of phospholipase C-γ (PLC-γ) isozymes. Like most other PLCs, PLC-γ1 is basally autoinhibited by its X-Y linker, which separates the X- and Y-boxes of the catalytic core. The C-terminal SH2 (cSH2) domain within the X-Y linker is the critical determinant for autoinhibition of phospholipase activity. Release of autoinhibition requires an intramolecular interaction between the cSH2 domain and a phosphorylated tyrosine, Tyr783, also located within the X-Y linker. The molecular mechanisms that mediate autoinhibition and phosphorylation-induced activation have not been defined. Here, we describe structures of the cSH2 domain both alone and bound to a PLC-γ1 peptide encompassing phosphorylated Tyr783. The cSH2 domain remains largely unaltered by peptide engagement. Point mutations in the cSH2 domain located at the interface with the peptide were sufficient to constitutively activate PLC-γ1, suggesting that peptide engagement directly interferes with the capacity of the cSH2 domain to block the lipase active site. This idea is supported by mutations in a complementary surface of the catalytic core that also enhanced phospholipase activity.
- Department of Pharmacology and ‡Department of Biochemistry and Biophysics and Lineberger Comprehensive Cancer Center, University of North Carolina School of Medicine , Chapel Hill, North Carolina 27599-7365, United States.
Organizational Affiliation: