Identification of Inhibitors of PvdQ, an Enzyme Involved in the Synthesis of the Siderophore Pyoverdine.
Wurst, J.M., Drake, E.J., Theriault, J.R., Jewett, I.T., VerPlank, L., Perez, J.R., Dandapani, S., Palmer, M., Moskowitz, S.M., Schreiber, S.L., Munoz, B., Gulick, A.M.(2014) ACS Chem Biol 9: 1536-1544
- PubMed: 24824984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/cb5001586
- Primary Citation Related Structures:
4K2F, 4K2G - PubMed Abstract:
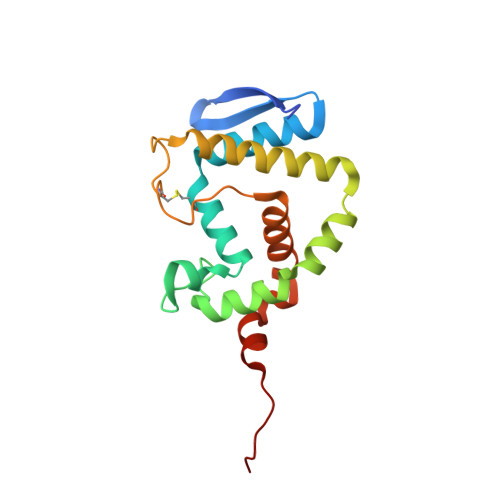
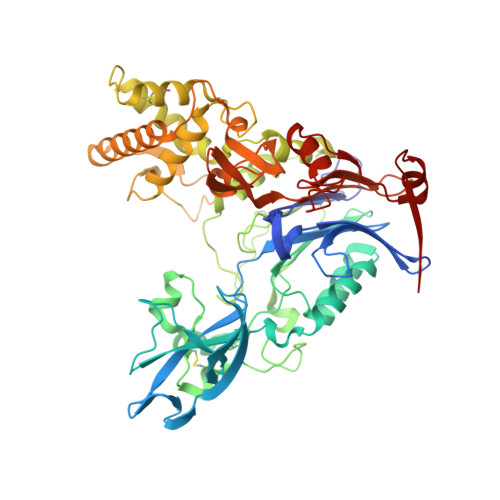
Pseudomonas aeruginosa produces the peptide siderophore pyoverdine, which is used to acquire essential Fe(3+) ions from the environment. PvdQ, an Ntn hydrolase, is required for the biosynthesis of pyoverdine. PvdQ knockout strains are not infectious in model systems, suggesting that disruption of siderophore production via PvdQ inhibition could be exploited as a target for novel antibacterial agents, by preventing cells from acquiring iron in the low iron environments of most biological settings. We have previously described a high-throughput screen to identify inhibitors of PvdQ that identified inhibitors with IC50 values of ∼100 μM. Here, we describe the discovery of ML318, a biaryl nitrile inhibitor of PvdQ acylase. ML318 inhibits PvdQ in vitro (IC50 = 20 nM) by binding in the acyl-binding site, as confirmed by the X-ray crystal structure of PvdQ bound to ML318. Additionally, the PvdQ inhibitor is active in a whole cell assay, preventing pyoverdine production and limiting the growth of P. aeruginosa under iron-limiting conditions.
- The Broad Institute , Cambridge, Massachusetts 02142, United States.
Organizational Affiliation: