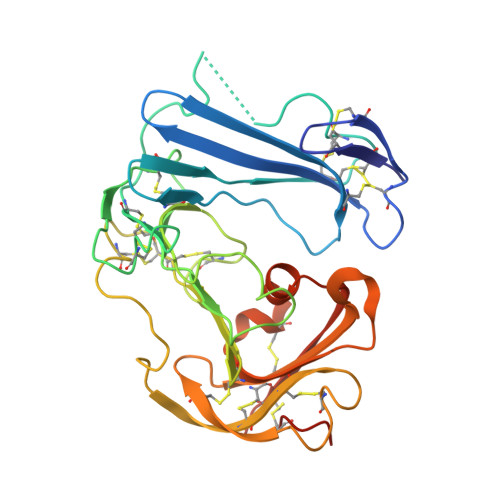
Identification of a New Epitope in uPAR as a Target for the Cancer Therapeutic Monoclonal Antibody ATN-658, a Structural Homolog of the uPAR Binding Integrin CD11b ( alpha M)
Xu, X., Cai, Y., Wei, Y., Donate, F., Juarez, J., Parry, G., Chen, L., Meehan, E.J., Ahn, R.W., Ugolkov, A., Dubrovskyi, O., O'halloran, T.V., Huang, M., Mazar, A.P.(2014) PLoS One 9: e85349-e85349
- PubMed: 24465541 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0085349
- Primary Citation Related Structures:
4K23, 4K24 - PubMed Abstract:
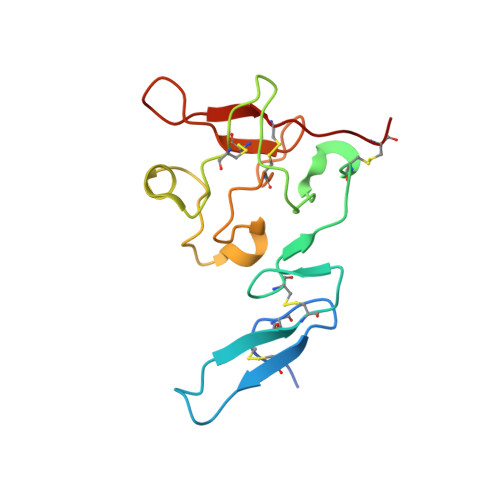
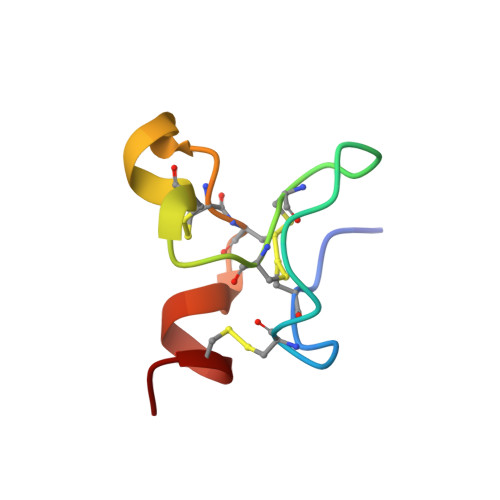
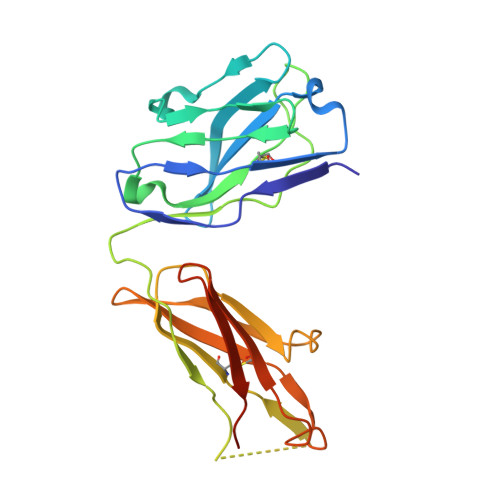
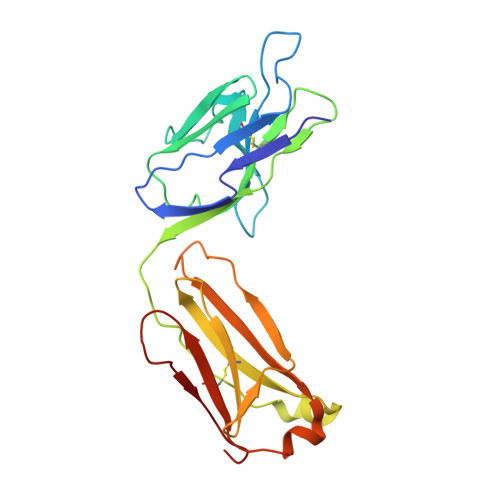
The urokinase plasminogen activator receptor (uPAR) plays a role in tumor progression and has been proposed as a target for the treatment of cancer. We recently described the development of a novel humanized monoclonal antibody that targets uPAR and has anti-tumor activity in multiple xenograft animal tumor models. This antibody, ATN-658, does not inhibit ligand binding (i.e. uPA and vitronectin) to uPAR and its mechanism of action remains unclear. As a first step in understanding the anti-tumor activity of ATN-658, we set out to identify the epitope on uPAR to which ATN-658 binds. Guided by comparisons between primate and human uPAR, epitope mapping studies were performed using several orthogonal techniques. Systematic site directed and alanine scanning mutagenesis identified the region of aa 268-275 of uPAR as the epitope for ATN-658. No known function has previously been attributed to this epitope Structural insights into epitope recognition were obtained from structural studies of the Fab fragment of ATN-658 bound to uPAR. The structure shows that the ATN-658 binds to the DIII domain of uPAR, close to the C-terminus of the receptor, corroborating the epitope mapping results. Intriguingly, when bound to uPAR, the complementarity determining region (CDR) regions of ATN-658 closely mimic the binding regions of the integrin CD11b (αM), a previously identified uPAR ligand thought to be involved in leukocyte rolling, migration and complement fixation with no known role in tumor progression of solid tumors. These studies reveal a new functional epitope on uPAR involved in tumor progression and demonstrate a previously unrecognized strategy for the therapeutic targeting of uPAR.
- Division of Hemostasis and Thrombosis, Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, Massachusetts, United States of America.
Organizational Affiliation: