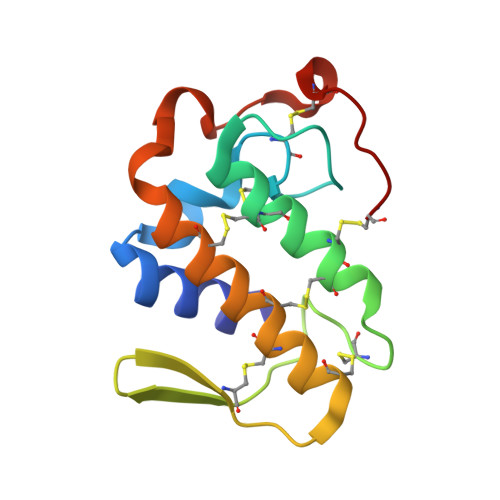
Structural bases for a complete myotoxic mechanism: Crystal structures of two non-catalytic phospholipases A2-like from Bothrops brazili venom.
Fernandes, C.A.H., Comparetti, E.J., Borges, R.J., Huancahuire-Vega, S., Ponce-Soto, L.A., Marangoni, S., Soares, A.M., Fontes, M.R.M.(2013) Biochim Biophys Acta 1834: 2772-2781
- PubMed: 24145104 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2013.10.009
- Primary Citation Related Structures:
4K06, 4K09 - PubMed Abstract:
Bothrops brazili is a snake found in the forests of the Amazonian region whose commercial therapeutic anti-bothropic serum has low efficacy for local myotoxic effects, resulting in an important public health problem in this area. Catalytically inactive phospholipases A2-like (Lys49-PLA2s) are among the main components from Bothrops genus venoms and are capable of causing drastic myonecrosis. Several studies have shown that the C-terminal region of these toxins, which includes a variable combination of positively charged and hydrophobic residues, is responsible for their activity. In this work we describe the crystal structures of two Lys49-PLA2s (BbTX-II and MTX-II) from B. brazili venom and a comprehensive structural comparison with several Lys49-PLA2s. Based on these results, two independent sites of interaction were identified between protein and membrane which leads to the proposition of a new myotoxic mechanism for bothropic Lys49-PLA2s composed of five different steps. This proposition is able to fully explain the action of these toxins and may be useful to develop efficient inhibitors to complement the conventional antivenom administration.
- Dep. de Física e Biofísica, Instituto de Biociências, UNESP - Universidade Estadual Paulista, Botucatu and Instituto Nacional de Ciência e Tecnologia em Toxinas, CNPq, Brazil.
Organizational Affiliation: