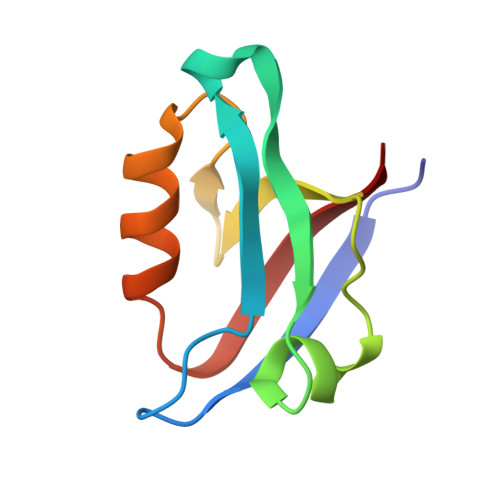
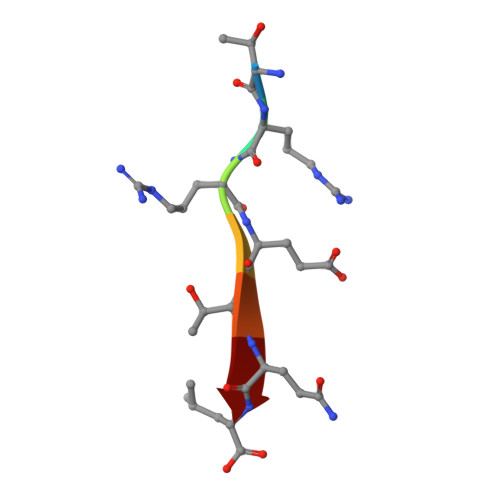
Stereochemical Preferences Modulate Affinity and Selectivity among Five PDZ Domains that Bind CFTR: Comparative Structural and Sequence Analyses.
Amacher, J.F., Cushing, P.R., Brooks, L., Boisguerin, P., Madden, D.R.(2014) Structure 22: 82-93
- PubMed: 24210758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.09.019
- Primary Citation Related Structures:
4JOE, 4JOF, 4JOG, 4JOH, 4JOJ, 4JOK, 4JOP, 4JOR, 4K6Y, 4K72, 4K75, 4K76, 4K78 - PubMed Abstract:
PDZ domain interactions are involved in signaling and trafficking pathways that coordinate crucial cellular processes. Alignment-based PDZ binding motifs identify the few most favorable residues at certain positions along the peptide backbone. However, sequences that bind the CAL (CFTR-associated ligand) PDZ domain reveal only a degenerate motif that overpredicts the true number of high-affinity interactors. Here, we combine extended peptide-array motif analysis with biochemical techniques to show that non-motif "modulator" residues influence CAL binding. The crystallographic structures of 13 CAL:peptide complexes reveal defined, but accommodating stereochemical environments at non-motif positions, which are reflected in modulator preferences uncovered by multisequence substitutional arrays. These preferences facilitate the identification of high-affinity CAL binding sequences and differentially affect CAL and NHERF PDZ binding. As a result, they also help determine the specificity of a PDZ domain network that regulates the trafficking of CFTR at the apical membrane.
- Department of Biochemistry, Geisel School of Medicine at Dartmouth, Hanover, NH 03755, USA.
Organizational Affiliation: