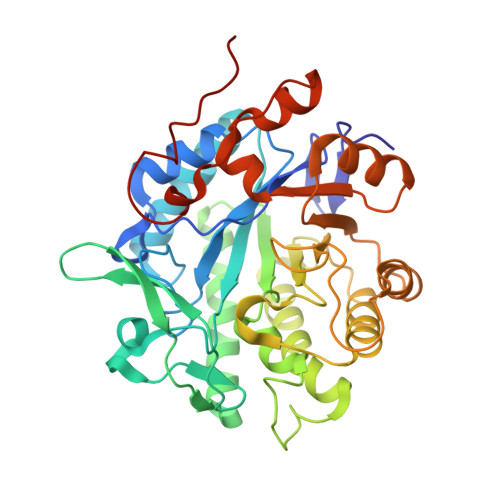
The Structure of Glycerol Trinitrate Reductase NerA from Agrobacterium radiobacter Reveals the Molecular Reason for Nitro- and Ene-Reductase Activity in OYE Homologues.
Oberdorfer, G., Binter, A., Wallner, S., Durchschein, K., Hall, M., Faber, K., Macheroux, P., Gruber, K.(2013) Chembiochem 14: 836-845
- PubMed: 23606302 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cbic.201300136
- Primary Citation Related Structures:
4JIC, 4JIP, 4JIQ - PubMed Abstract:
In recent years, Old Yellow Enzymes (OYEs) and their homologues have found broad application in the efficient asymmetric hydrogenation of activated C=C bonds with high selectivities and yields. Members of this class of enzymes have been found in many different organisms and are rather diverse on the sequence level, with pairwise identities as low as 20 %, but they exhibit significant structural similarities with the adoption of a conserved (αβ)(8)-barrel fold. Some OYEs have been shown not only to reduce C=C double bonds, but also to be capable of reducing nitro groups in both saturated and unsaturated substrates. In order to understand this dual activity we determined and analyzed X-ray crystal structures of NerA from Agrobacterium radiobacter, both in its apo form and in complex with 4-hydroxybenzaldehyde and with 1-nitro-2-phenylpropene. These structures, together with spectroscopic studies of substrate binding to several OYEs, indicate that nitro-containing substrates can bind to OYEs in different binding modes, one of which leads to C=C double bond reduction and the other to nitro group reduction.
- ACIB--Austrian Centre of Industrial Biotechnology, Petergasse 14, 8010 Graz, Austria.
Organizational Affiliation: