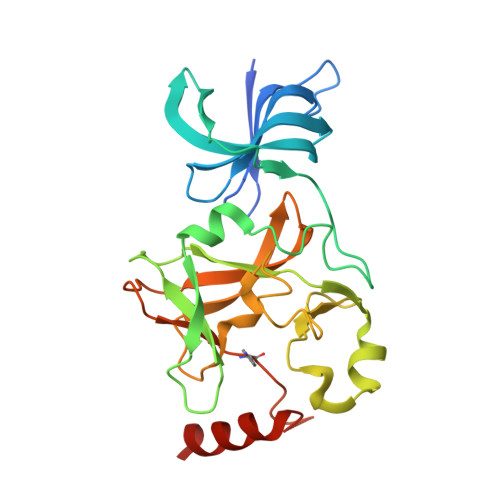
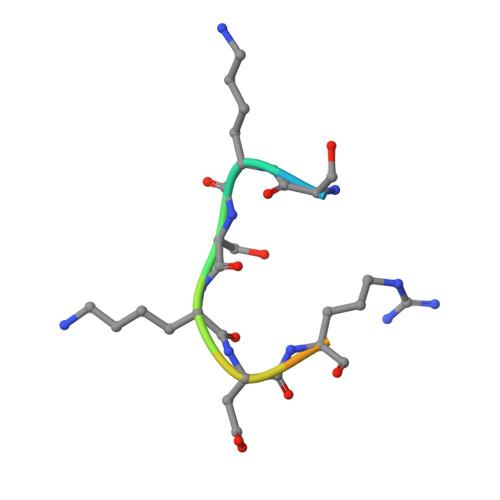
Methyl CH O Hydrogen Bonds Orchestrate AdoMet-Dependent Methylation
Horowitz, S., Dirk, L.M.A., Yesselman, J.D., Nimtz, J.S., Del Rizzo, P.A., Vander Meulen, K.A., Butcher, S.E., Mehl, R.A., Houtz, R.L., Al-Hashimi, H.M., Trievel, R.C.To be published.