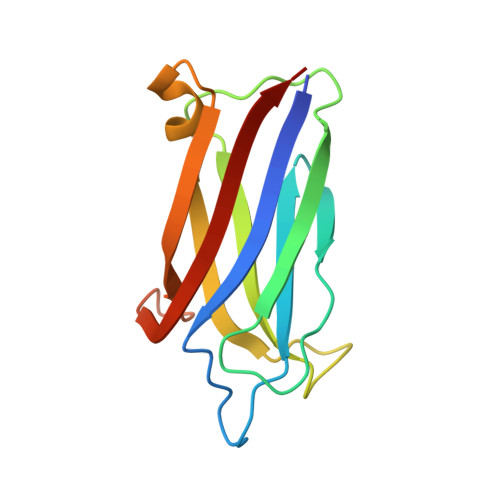
Alternate Splicing of Dysferlin C2A Confers Ca(2+)-Dependent and Ca(2+)-Independent Binding for Membrane Repair.
Fuson, K., Rice, A., Mahling, R., Snow, A., Nayak, K., Shanbhogue, P., Meyer, A.G., Redpath, G.M., Hinderliter, A., Cooper, S.T., Sutton, R.B.(2014) Structure 22: 104-115
- PubMed: 24239457
- DOI: https://doi.org/10.1016/j.str.2013.10.001
- Primary Citation of Related Structures:
4IHB, 4IQH - PubMed Abstract:
Dysferlin plays a critical role in the Ca²⁺-dependent repair of microlesions that occur in the muscle sarcolemma. Of the seven C2 domains in dysferlin, only C2A is reported to bind both Ca²⁺ and phospholipid, thus acting as a key sensor in membrane repair. Dysferlin C2A exists as two isoforms, the "canonical" C2A and C2A variant 1 (C2Av1). Interestingly, these isoforms have markedly different responses to Ca²⁺ and phospholipid. Structural and thermodynamic analyses are consistent with the canonical C2A domain as a Ca²⁺-dependent, phospholipid-binding domain, whereas C2Av1 would likely be Ca²⁺-independent under physiological conditions. Additionally, both isoforms display remarkably low free energies of stability, indicative of a highly flexible structure. The inverted ligand preference and flexibility for both C2A isoforms suggest the capability for both constitutive and Ca²⁺-regulated effector interactions, an activity that would be essential in its role as a mediator of membrane repair.
- Department of Cell Physiology and Molecular Biophysics, Texas Tech University Health Sciences Center, Lubbock, TX 79430, USA; Center for Membrane Protein Research, Texas Tech University Health Sciences Center, Lubbock, TX 79430, USA.
Organizational Affiliation: