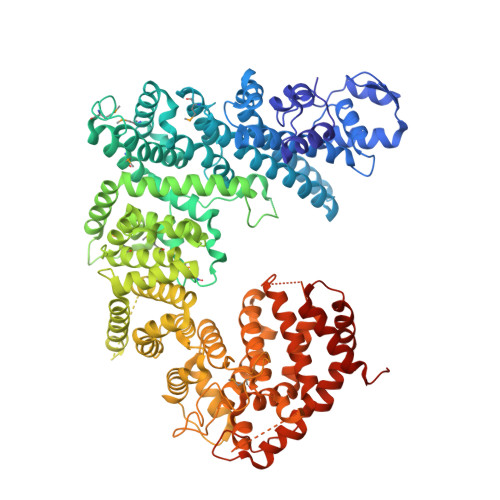
Structure, dynamics, evolution, and function of a major scaffold component in the nuclear pore complex.
Sampathkumar, P., Kim, S.J., Upla, P., Rice, W.J., Phillips, J., Timney, B.L., Pieper, U., Bonanno, J.B., Fernandez-Martinez, J., Hakhverdyan, Z., Ketaren, N.E., Matsui, T., Weiss, T.M., Stokes, D.L., Sauder, J.M., Burley, S.K., Sali, A., Rout, M.P., Almo, S.C.(2013) Structure 21: 560-571
- PubMed: 23499021
- DOI: https://doi.org/10.1016/j.str.2013.02.005
- Primary Citation Related Structures:
4IFQ - PubMed Abstract:
The nuclear pore complex, composed of proteins termed nucleoporins (Nups), is responsible for nucleocytoplasmic transport in eukaryotes. Nuclear pore complexes (NPCs) form an annular structure composed of the nuclear ring, cytoplasmic ring, a membrane ring, and two inner rings. Nup192 is a major component of the NPC's inner ring. We report the crystal structure of Saccharomyces cerevisiae Nup192 residues 2-960 [ScNup192(2-960)], which adopts an α-helical fold with three domains (i.e., D1, D2, and D3). Small angle X-ray scattering and electron microscopy (EM) studies reveal that ScNup192(2-960) could undergo long-range transition between "open" and "closed" conformations. We obtained a structural model of full-length ScNup192 based on EM, the structure of ScNup192(2-960), and homology modeling. Evolutionary analyses using the ScNup192(2-960) structure suggest that NPCs and vesicle-coating complexes are descended from a common membrane-coating ancestral complex. We show that suppression of Nup192 expression leads to compromised nuclear transport and hypothesize a role for Nup192 in modulating the permeability of the NPC central channel.
- Department of Biochemistry, Ullmann Building, Room 409, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA.
Organizational Affiliation: