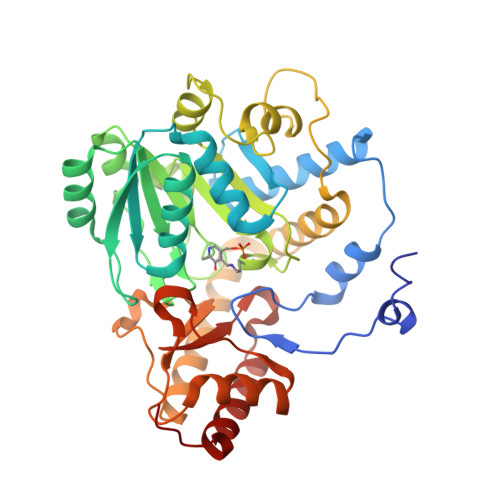
Crystal structure of the S187F variant of human liver alanine: Aminotransferase associated with primary hyperoxaluria type I and its functional implications.
Oppici, E., Fodor, K., Paiardini, A., Williams, C., Voltattorni, C.B., Wilmanns, M., Cellini, B.(2013) Proteins 81: 1457-1465
- PubMed: 23589421 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24300
- Primary Citation Related Structures:
4I8A - PubMed Abstract:
The substitution of Ser187, a residue located far from the active site of human liver peroxisomal alanine:glyoxylate aminotransferase (AGT), by Phe gives rise to a variant associated with primary hyperoxaluria type I. Unexpectedly, previous studies revealed that the recombinant form of S187F exhibits a remarkable loss of catalytic activity, an increased pyridoxal 5'-phosphate (PLP) binding affinity and a different coenzyme binding mode compared with normal AGT. To shed light on the structural elements responsible for these defects, we solved the crystal structure of the variant to a resolution of 2.9 Å. Although the overall conformation of the variant is similar to that of normal AGT, we noticed: (i) a displacement of the PLP-binding Lys209 and Val185, located on the re and si side of PLP, respectively, and (ii) slight conformational changes of other active site residues, in particular Trp108, the base stacking residue with the pyridine cofactor moiety. This active site perturbation results in a mispositioning of the AGT-pyridoxamine 5'-phosphate (PMP) complex and of the external aldimine, as predicted by molecular modeling studies. Taken together, both predicted and observed movements caused by the S187F mutation are consistent with the following functional properties of the variant: (i) a 300- to 500-fold decrease in both the rate constant of L-alanine half-transamination and the kcat of the overall transamination, (ii) a different PMP binding mode and affinity, and (iii) a different microenvironment of the external aldimine. Proposals for the treatment of patients bearing S187F mutation are discussed on the basis of these results.
- Department of Life Sciences and Reproduction, Section of Biological Chemistry, University of Verona, Verona, Italy.
Organizational Affiliation: