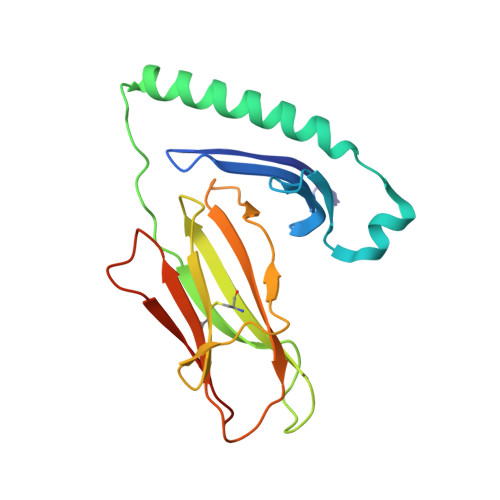
Disruption of hydrogen bonds between major histocompatibility complex class II and the peptide N-terminus is not sufficient to form a human leukocyte antigen-DM receptive state of major histocompatibility complex class II.
Schulze, M.S., Anders, A.K., Sethi, D.K., Call, M.J.(2013) PLoS One 8: e69228-e69228
- PubMed: 23976922 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0069228
- Primary Citation Related Structures:
4I5B - PubMed Abstract:
Peptide presentation by MHC class II is of critical importance to the function of CD4+ T cells. HLA-DM resides in the endosomal pathway and edits the peptide repertoire of newly synthesized MHC class II molecules before they are exported to the cell surface. HLA-DM ensures MHC class II molecules bind high affinity peptides by targeting unstable MHC class II:peptide complexes for peptide exchange. Research over the past decade has implicated the peptide N-terminus in modulating the ability of HLA-DM to target a given MHC class II:peptide combination. In particular, attention has been focused on both the hydrogen bonds between MHC class II and peptide, and the occupancy of the P1 anchor pocket. We sought to solve the crystal structure of a HLA-DR1 molecule containing a truncated hemagglutinin peptide missing three N-terminal residues compared to the full-length sequence (residues 306-318) to determine the nature of the MHC class II:peptide species that binds HLA-DM. Here we present structural evidence that HLA-DR1 that is loaded with a peptide truncated to the P1 anchor residue such that it cannot make select hydrogen bonds with the peptide N-terminus, adopts the same conformation as molecules loaded with full-length peptide. HLA-DR1:peptide combinations that were unable to engage up to four key hydrogen bonds were also unable to bind HLA-DM, while those truncated to the P2 residue bound well. These results indicate that the conformational changes in MHC class II molecules that are recognized by HLA-DM occur after disengagement of the P1 anchor residue.
- Department of Cancer Immunology & AIDS, Dana-Farber Cancer Institute, Boston, Massachusetts, United States of America.
Organizational Affiliation: