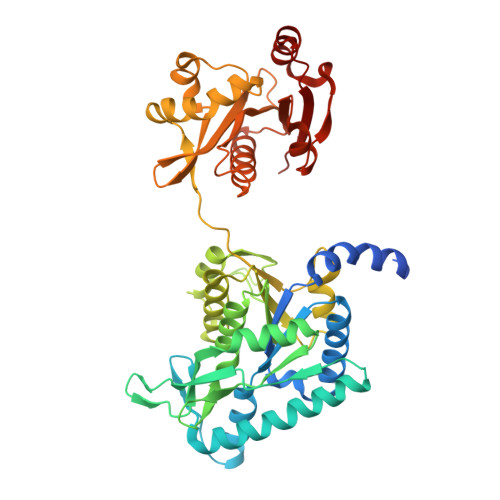
Structure of the complex of Neisseria gonorrhoeae N-acetyl-l-glutamate synthase with a bound bisubstrate analog.
Zhao, G., Allewell, N.M., Tuchman, M., Shi, D.(2013) Biochem Biophys Res Commun 430: 1253-1258
- PubMed: 23261468 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrc.2012.12.064
- Primary Citation Related Structures:
4I49 - PubMed Abstract:
N-Acetyl-L-glutamate synthase catalyzes the conversion of AcCoA and glutamate to CoA and N-acetyl-L-glutamate (NAG), the first step of the arginine biosynthetic pathway in lower organisms. In mammals, NAG is an obligate cofactor of carbamoyl phosphate synthetase I in the urea cycle. We have previously reported the structures of NAGS from Neisseria gonorrhoeae (ngNAGS) with various substrates bound. Here we reported the preparation of the bisubstrate analog, CoA-S-acetyl-L-glutamate, the crystal structure of ngNAGS with CoA-NAG bound, and kinetic studies of several active site mutants. The results are consistent with a one-step nucleophilic addition-elimination mechanism with Glu353 as the catalytic base and Ser392 as the catalytic acid. The structure of the ngNAGS-bisubstrate complex together with the previous ngNAGS structures delineates the catalytic reaction path for ngNAGS.
- Center for Genetic Medicine Research and Department of Integrative Systems Biology, Children's National Medical Center, The George Washington University, Washington, DC 20010, USA.
Organizational Affiliation: