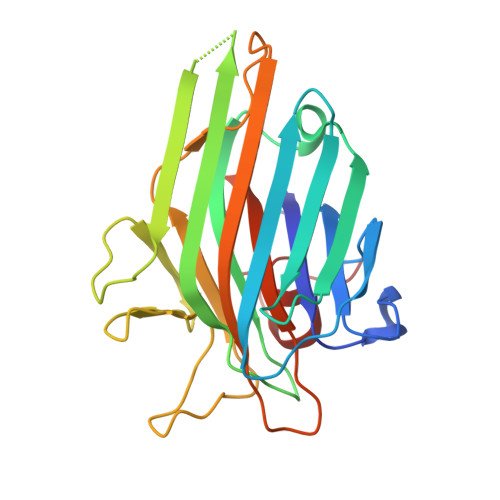
Crystal structure of Canavalia maritima seeds lectin (ConM) co-crystalized with gamma-aminobutyric acid (GABA) and soaked with adenine
Delatorre, P., Silva-Filho, J.C., Nobrega, R.B., Gadelha, C.A.A., Cavada, B.S., Rocha, B.A.M., Santi-Gadelha, T., Teixeira, C.S.To be published.