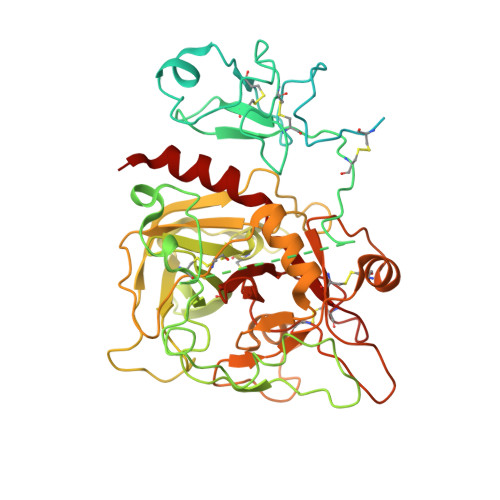
Crystal structure of prothrombin reveals conformational flexibility and mechanism of activation.
Pozzi, N., Chen, Z., Gohara, D.W., Niu, W., Heyduk, T., Di Cera, E.(2013) J Biological Chem 288: 22734-22744
- PubMed: 23775088 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.466946
- Primary Citation Related Structures:
4HZH - PubMed Abstract:
The zymogen prothrombin is composed of fragment 1 containing a Gla domain and kringle-1, fragment 2 containing kringle-2, and a protease domain containing A and B chains. The prothrombinase complex assembled on the surface of platelets converts prothrombin to thrombin by cleaving at Arg-271 and Arg-320. The three-dimensional architecture of prothrombin and the molecular basis of its activation remain elusive. Here we report the first x-ray crystal structure of prothrombin as a Gla-domainless construct carrying an Ala replacement of the catalytic Ser-525. Prothrombin features a conformation 80 Å long, with fragment 1 positioned at a 36° angle relative to the main axis of fragment 2 coaxial to the protease domain. High flexibility of the linker connecting the two kringles suggests multiple arrangements for kringle-1 relative to the rest of the prothrombin molecule. Luminescence resonance energy transfer measurements detect two distinct conformations of prothrombin in solution, in a 3:2 ratio, with the distance between the two kringles either fully extended (54 ± 2 Å) or partially collapsed (≤34 Å) as seen in the crystal structure. A molecular mechanism of prothrombin activation emerges from the structure. Of the two sites of cleavage, Arg-271 is located in a disordered region connecting kringle-2 to the A chain, but Arg-320 is well defined within the activation domain and is not accessible to proteolysis in solution. Burial of Arg-320 prevents prothrombin autoactivation and directs prothrombinase to cleave at Arg-271 first. Reversal of the local electrostatic potential then redirects prothrombinase toward Arg-320, leading to thrombin generation via the prethrombin-2 intermediate.
- Edward A. Doisy Department of Biochemistry and Molecular Biology, Saint Louis University School of Medicine, St. Louis, Missouri 63104, USA.
Organizational Affiliation: