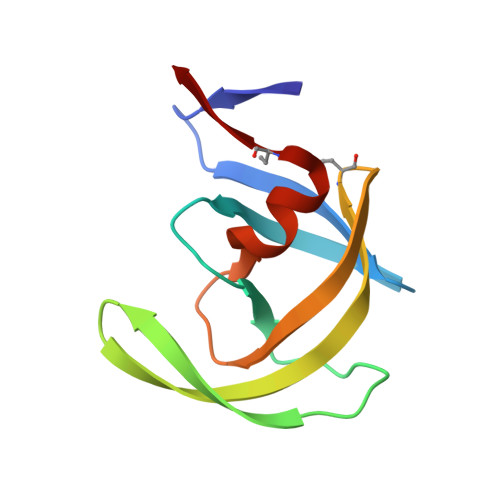
Structure of complex of synthetic HIV-1 protease with a substrate-based inhibitor at 2.3 A resolution.
Miller, M., Schneider, J., Sathyanarayana, B.K., Toth, M.V., Marshall, G.R., Clawson, L., Selk, L., Kent, S.B., Wlodawer, A.(1989) Science 246: 1149-1152
- PubMed: 2686029 Search on PubMed
- DOI: https://doi.org/10.1126/science.2686029
- Primary Citation Related Structures:
4HVP - PubMed Abstract:
The structure of a complex between a peptide inhibitor with the sequence N-acetyl-Thr-Ile-Nle-psi[CH2-NH]-Nle-Gln-Arg.amide (Nle, norleucine) with chemically synthesized HIV-1 (human immunodeficiency virus 1) protease was determined at 2.3 A resolution (R factor of 0.176). Despite the symmetric nature of the unliganded enzyme, the asymmetric inhibitor lies in a single orientation and makes extensive interactions at the interface between the two subunits of the homodimeric protein. Compared with the unliganded enzyme, the protein molecule underwent substantial changes, particularly in an extended region corresponding to the "flaps" (residues 35 to 57 in each chain), where backbone movements as large as 7 A are observed.
- NCI-Frederick Cancer Research Facility, BRI-Basic Research Program, MD 21701.
Organizational Affiliation: