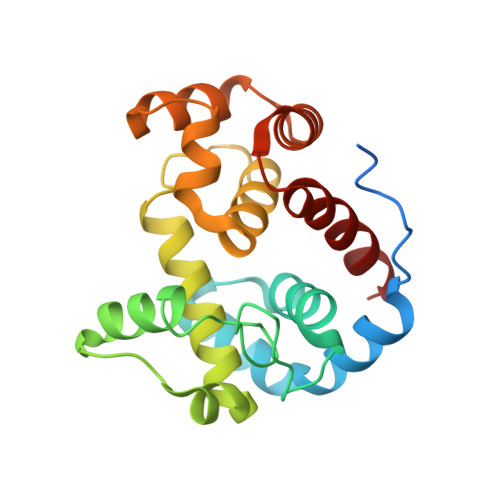
On the Mechanism of Peptidoglycan Binding and Cleavage by the endo-Specific Lytic Transglycosylase MltE from Escherichia coli.
Fibriansah, G., Gliubich, F.I., Thunnissen, A.M.(2012) Biochemistry 51: 9164-9177
- PubMed: 23075328 Search on PubMed
- DOI: https://doi.org/10.1021/bi300900t
- Primary Citation Related Structures:
3T36, 4HJV, 4HJY, 4HJZ - PubMed Abstract:
The lytic transglycosylase MltE from Escherichia coli is a periplasmic, outer membrane-attached enzyme that cleaves the β-1,4-glycosidic bonds between N-acetylmuramic acid and N-acetylglucosamine residues in the cell wall peptidoglycan, producing 1,6-anhydromuropeptides. Here we report three crystal structures of MltE: in a substrate-free state, in a binary complex with chitopentaose, and in a ternary complex with the glycopeptide inhibitor bulgecin A and the murodipeptide N-acetylglucosaminyl-N-acetylmuramyl-l-Ala-d-Glu. The substrate-bound structures allowed a detailed analysis of the saccharide-binding interactions in six subsites of the peptidoglycan-binding groove (subsites -4 to +2) and, combined with site-directed mutagenesis analysis, confirmed the role of Glu64 as catalytic acid/base. The structures permitted the precise modeling of a short glycan strand of eight saccharide residues, providing evidence for two additional subsites (+3 and +4) and revealing the productive conformational state of the substrate at subsites -1 and +1, where the glycosidic bond is cleaved. Full accessibility of the peptidoglycan-binding groove and preferential binding of an N-acetylmuramic acid residue in a (4)C(1) chair conformation at subsite +2 explain why MltE shows only endo- and no exo-specific activity toward glycan strands. The results further indicate that catalysis of glycosidic bond cleavage by MltE proceeds via distortion toward a sofa-like conformation of the N-acetylmuramic acid sugar ring at subsite -1 and by anchimeric assistance of the sugar's N-acetyl group, as shown previously for the lytic transglycosylases Slt70 and MltB.
- Laboratory of Biophysical Chemistry, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Nijenborgh 7, 9747 AG Groningen, The Netherlands.
Organizational Affiliation: