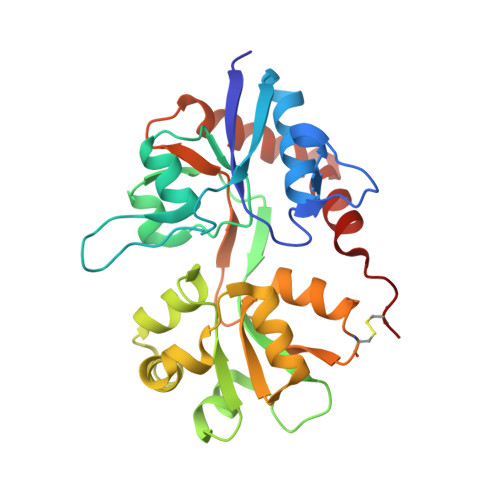
A photoswitchable neurotransmitter analogue bound to its receptor.
Reiter, A., Skerra, A., Trauner, D., Schiefner, A.(2013) Biochemistry 52: 8972-8974
- PubMed: 24295282 Search on PubMed
- DOI: https://doi.org/10.1021/bi4014402
- Primary Citation Related Structures:
4H8I - PubMed Abstract:
Incorporation of the azobenzene derivative gluazo, a synthetic photochromic ligand, into a kainate receptor allows for the optical control of neuronal activity. The crystal structure of gluazo bound to a dimeric GluK2 ligand-binding domain reveals one monomer in a closed conformation, occupied by gluazo, and the other in an open conformation, with a bound buffer molecule. The glutamate group of gluazo interacts like the natural glutamate ligand, while its trans-azobenzene moiety protrudes into a tunnel. This elongated cavity presumably cannot accommodate a cis-azobenzene, which explains the reversible activation of the receptor upon photoisomerization.
- Munich Center for Integrated Protein Science (CIPS-M).
Organizational Affiliation: